| Home | Papers | Reports | Projects | Code Fragments | Dissertations | Presentations | Posters | Proposals | Lectures given | Course notes |
Denoising of MALDI-TOF Mass Spectra
Werner Van Belle1* - werner@yellowcouch.org, werner.van.belle@gmail.com
Olav Mjaavatten2
1- Bioinformatics Group
Norut IT; Research Park; 9294 Tromsø; Norway
2- Proteomic Unit (Probe)
University of Bergen
; Bergen; Norway
* Corresponding author
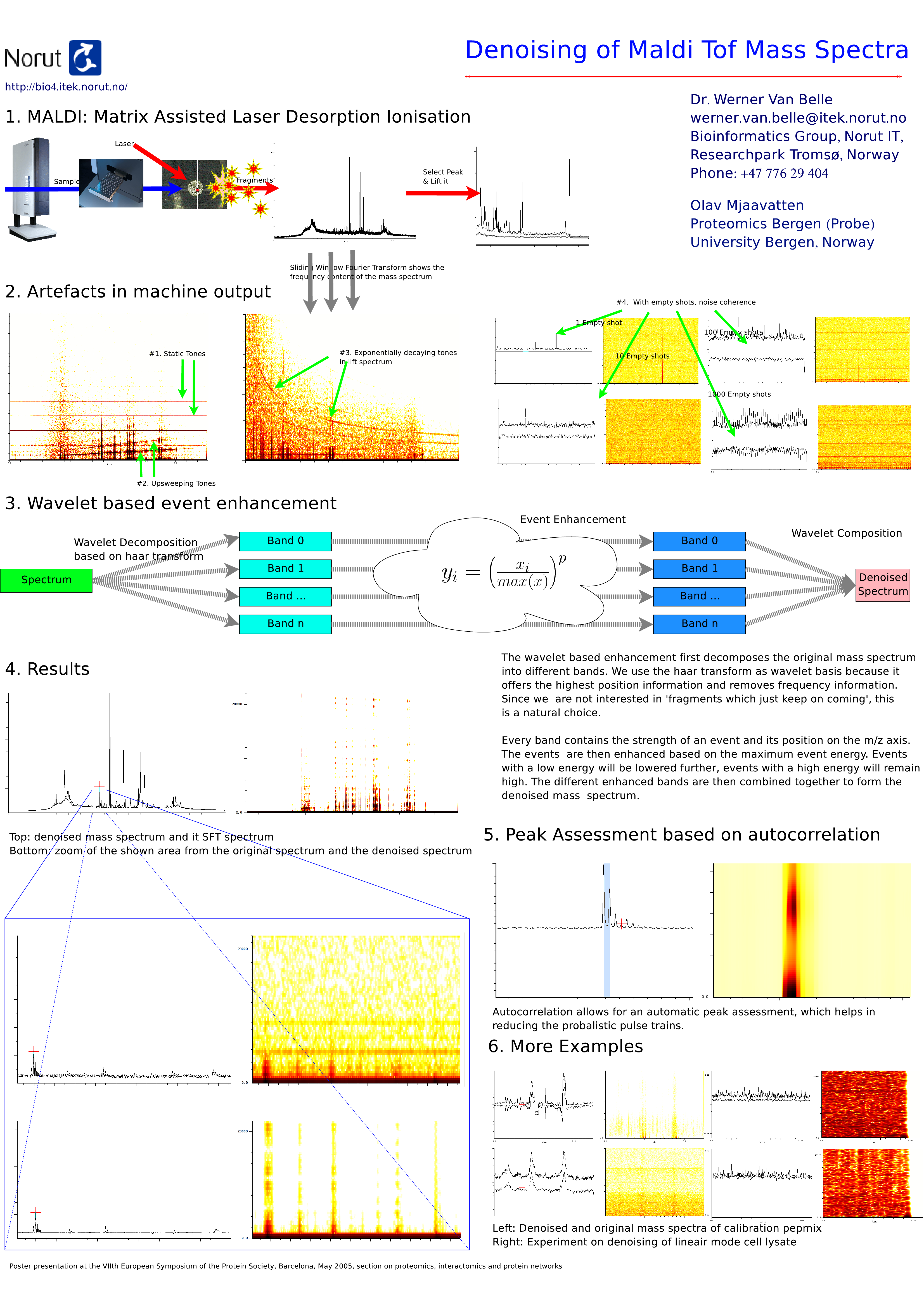
Abstract : MALDI-TOF mass spectrometry is a well known and widely used technique to fingerprint and sequence proteins. However widely used, the signal output of these machines often contain disturbing artefacts such as static tones, linearly up-sweeping tones, exponentially decaying tones and probabilistic pulse trains. Their presence reduces the accuracy of peak localization and might introduce phantom peaks. This further complicates a) automatic selection of peaks, b) makes it impossible to normalize mass spectrograms and c) in general reduces the performance of high throughput proteomics. We present a) a number of specialized algorithms to remove these artefacts, based on wavelets, and b) a number of techniques to automatically assign confidence scores to peaks, based on autocorrelation techniques and machine modelling. The combination of the presented techniques allow for a very accurate and automated sample analysis, which in turn helps in the determination of the sample content. Static tones are sine waves embedded in the signal, which can shift peak locations backward or forward over the mass/charge axis. Up-sweeping tones and decaying tones are sine waves of which the frequency goes linearly up, or exponentially down over the mass/charge axis, resulting in an immediate correlation between the accuracy of a peak-position and its position on the mass/charge axis. Probabilistic pulse trains are small pulses which might occur at specific positions on the mass/charge axis. Depending on the number of shots summed into one measurement different misinterpretations are possible. Either too few shots are taken, resulting in phantom peaks, or too many shots are taken in which the pulses will overrun the actual measurement
Keywords:
Reference:
Werner Van Belle, Olav Mjaavatten; Denoising of MALDI-TOF Mass Spectra; Presented at AOPC2005, The VIIth European Symposium of the Protein Society, section proteomics, interactomics and protein networks, Barcelona; Spain; May 2005
| http://werner.yellowcouch.org/ werner@yellowcouch.org |  |