| Home | Papers | Reports | Projects | Code Fragments | Dissertations | Presentations | Posters | Proposals | Lectures given | Course notes |
Microarrays, Protein Interaction Maps and Elevated Mazes
Werner Van Belle1* - werner@yellowcouch.org, werner.van.belle@gmail.com
1- Department of Medical Genetics (MEDGEN); Northern Norwegian Hospital; Tromsø; Norway
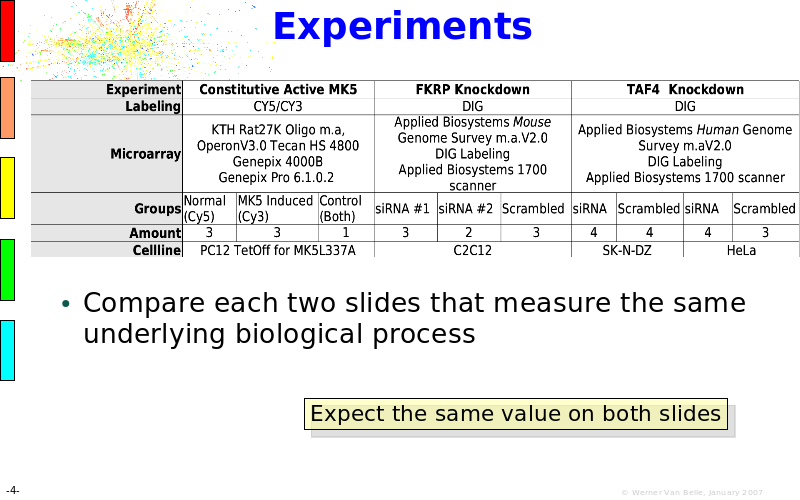
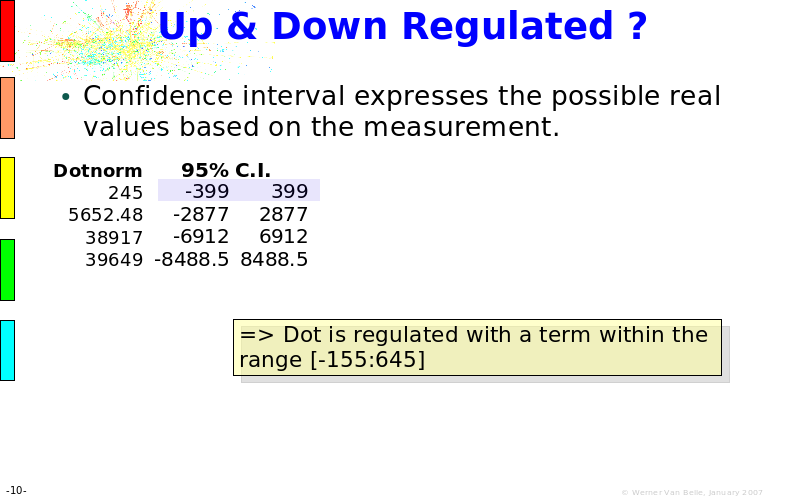
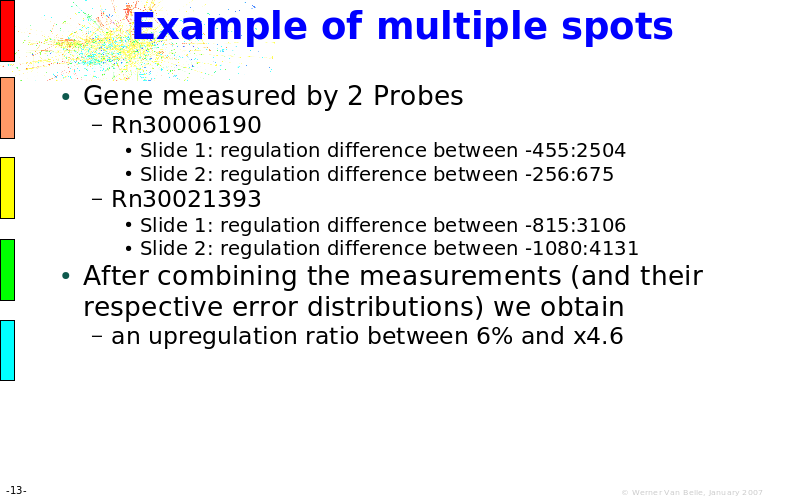
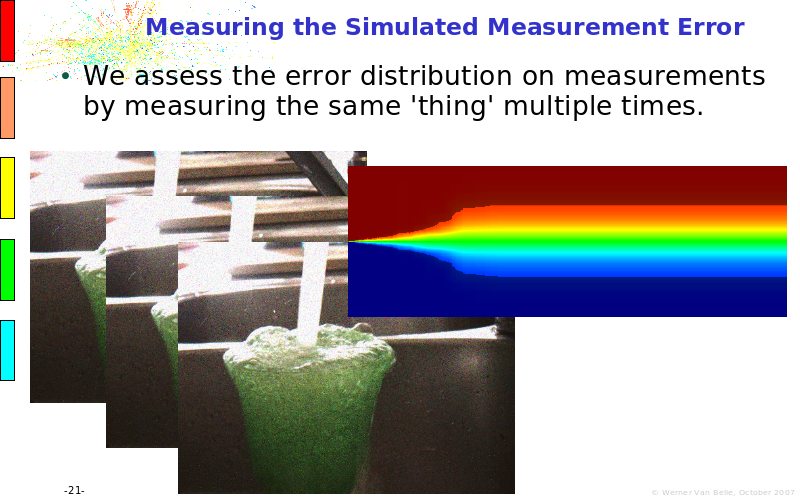
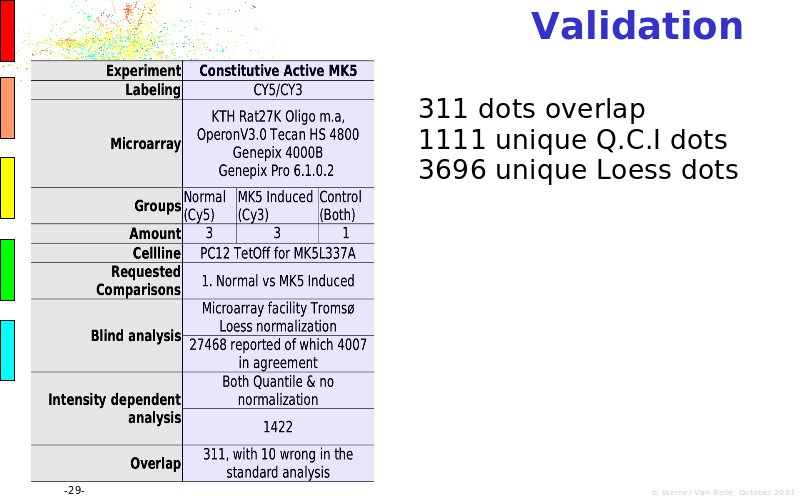
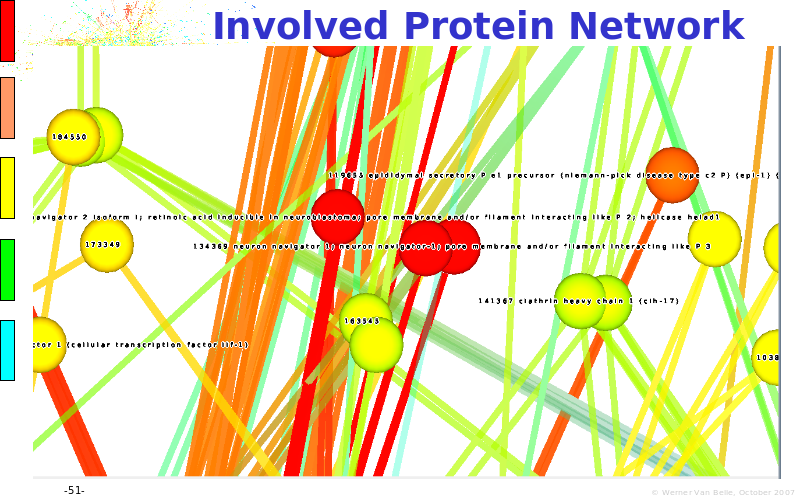
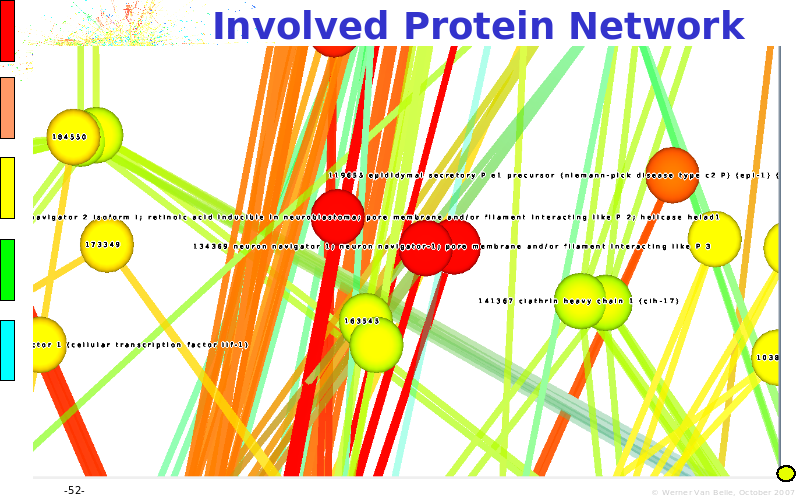
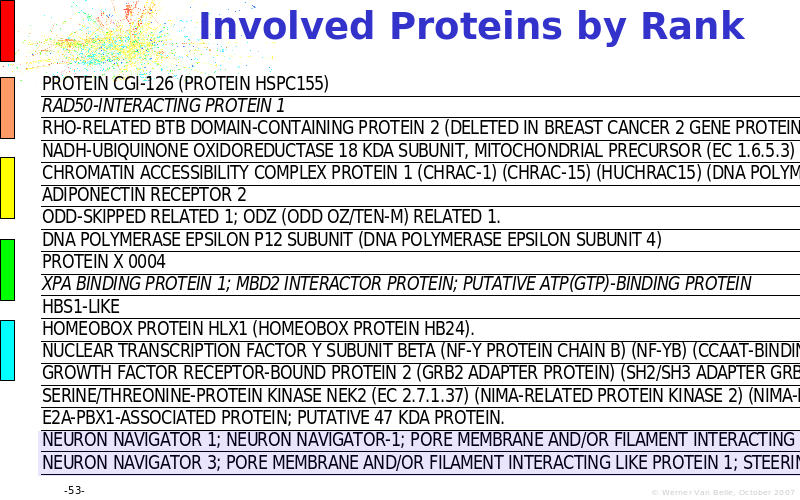
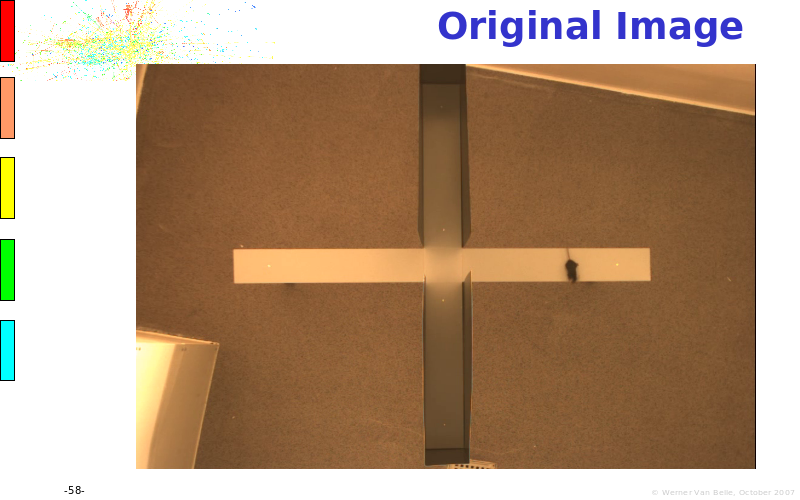
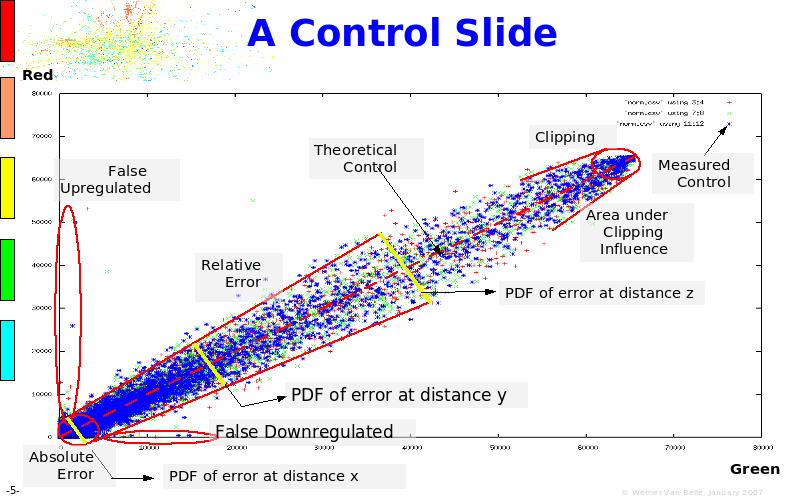
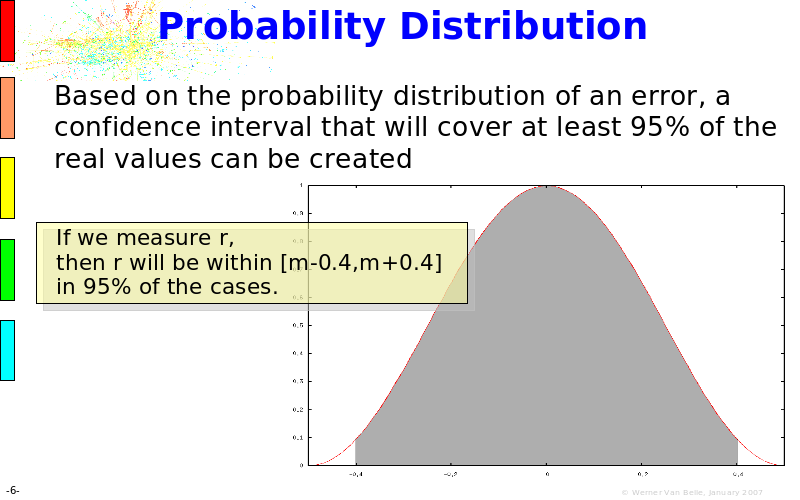
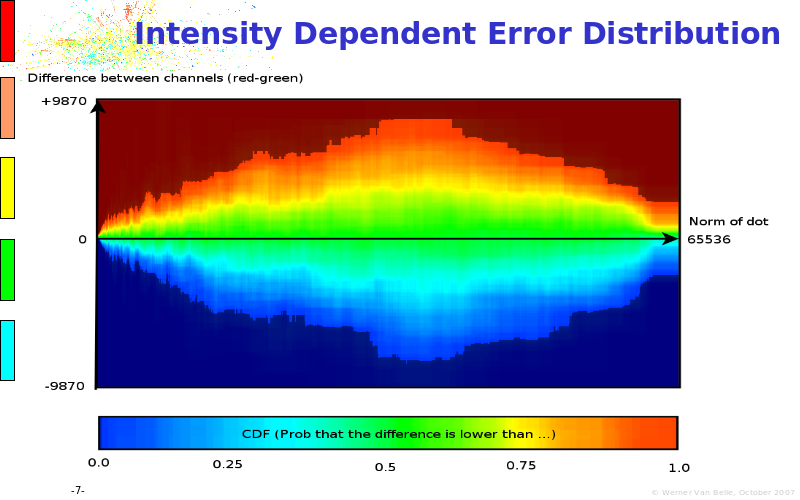
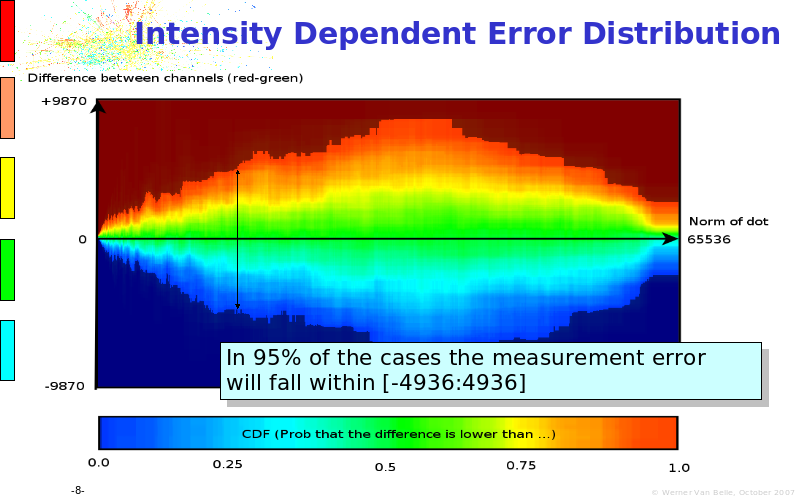
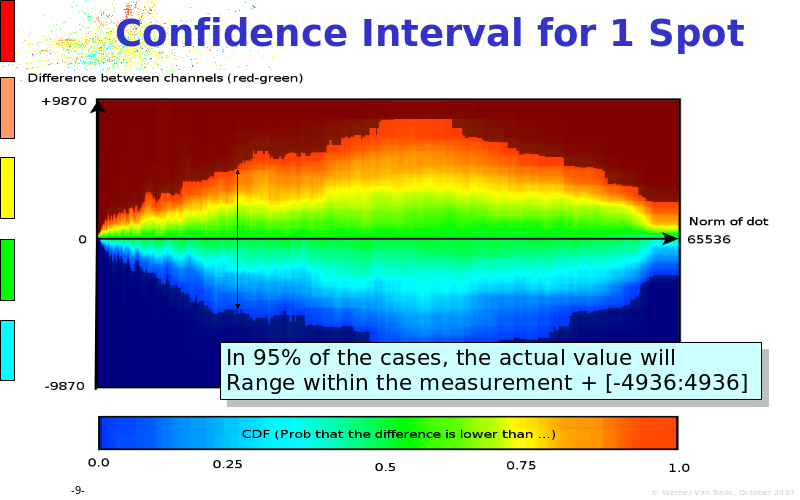
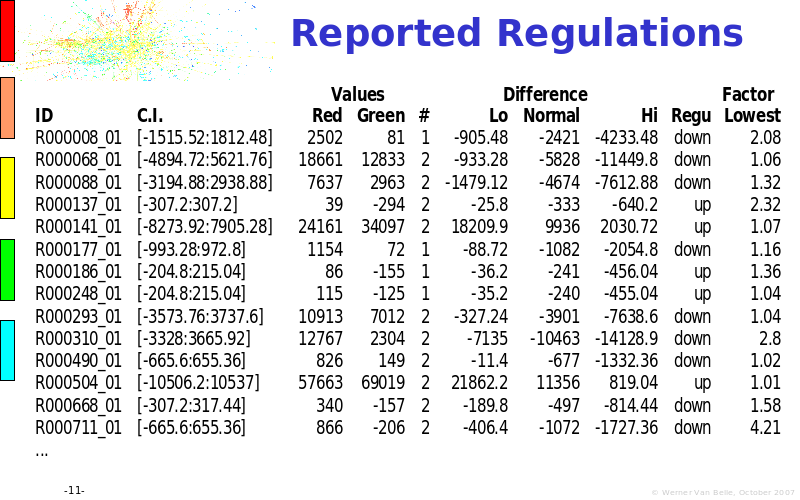
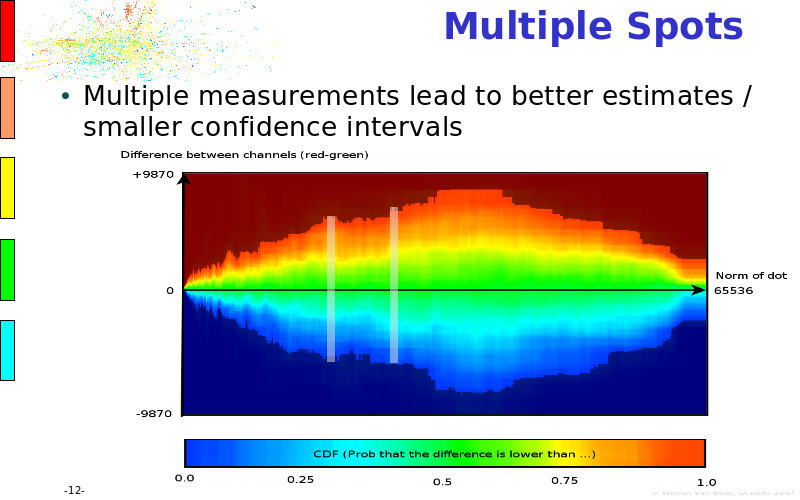
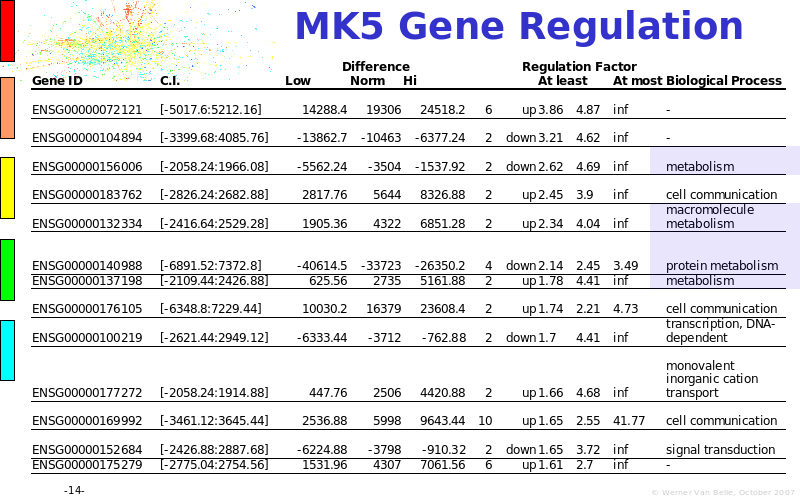
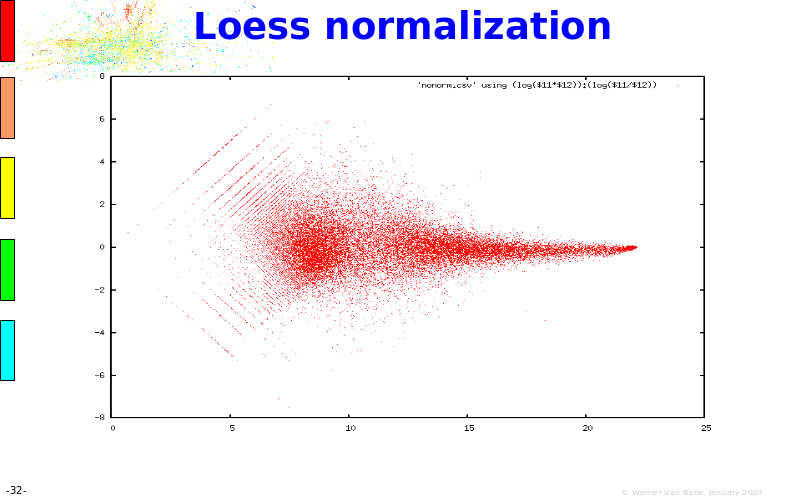
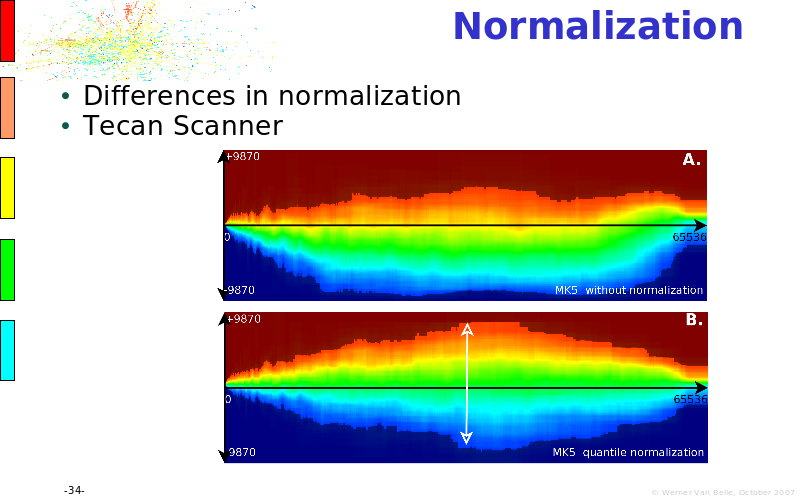
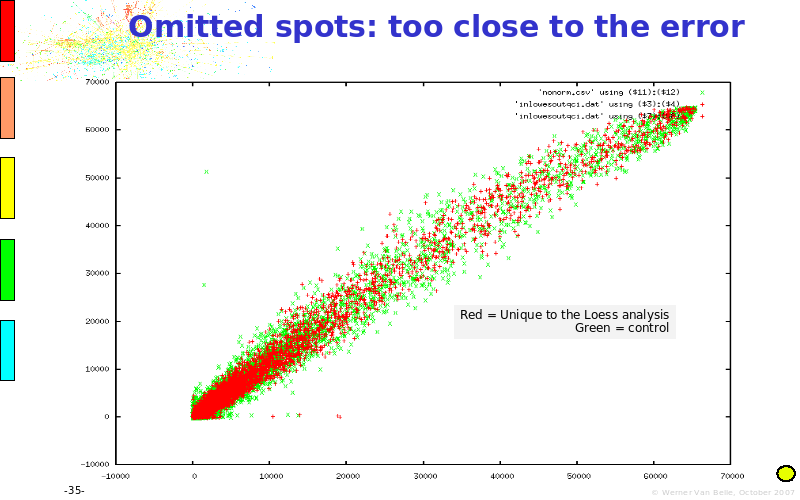
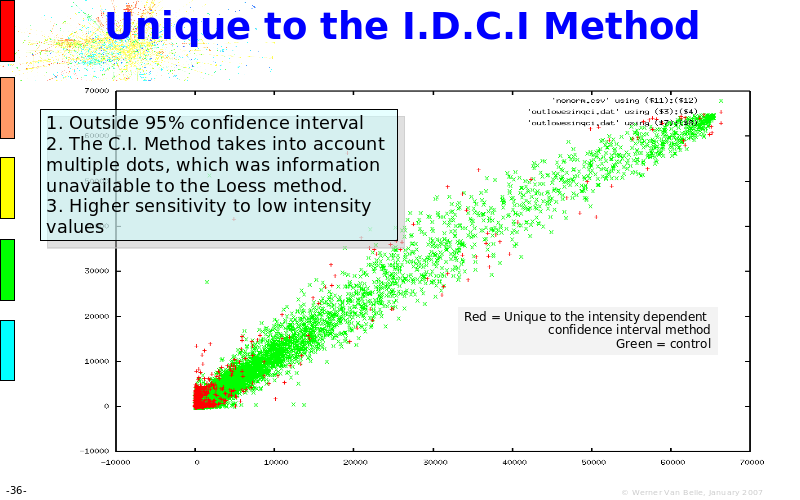
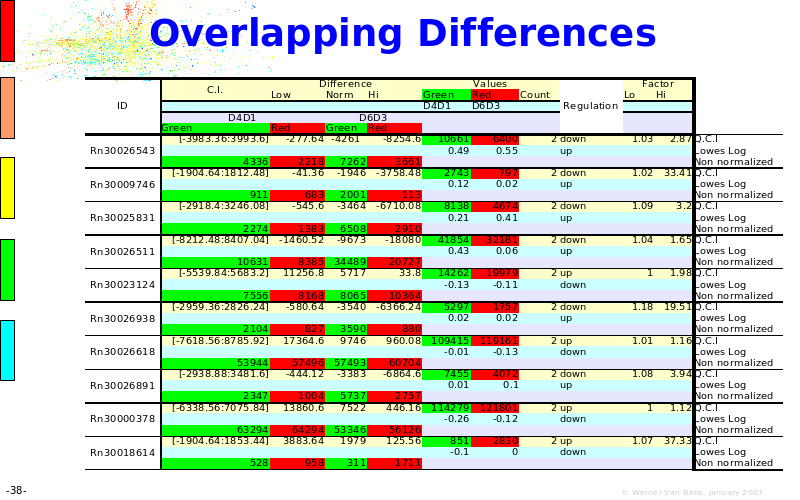
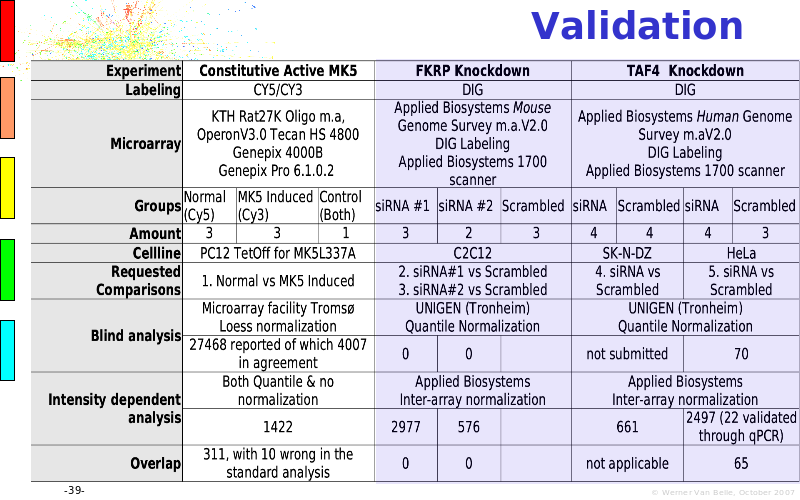
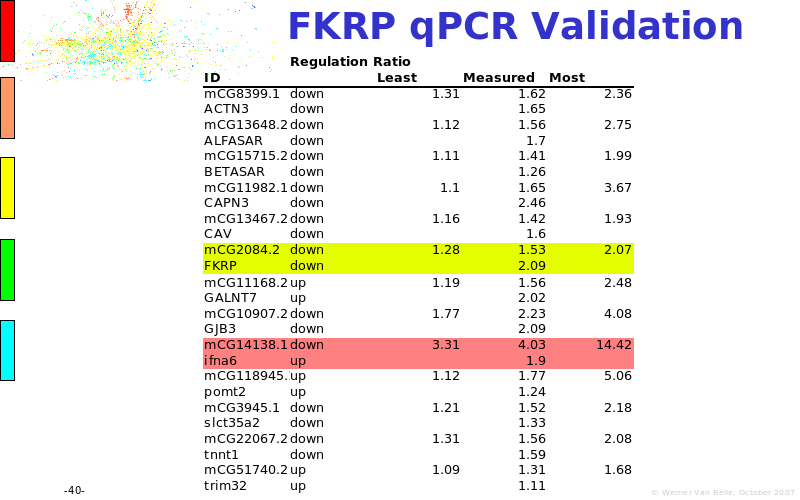
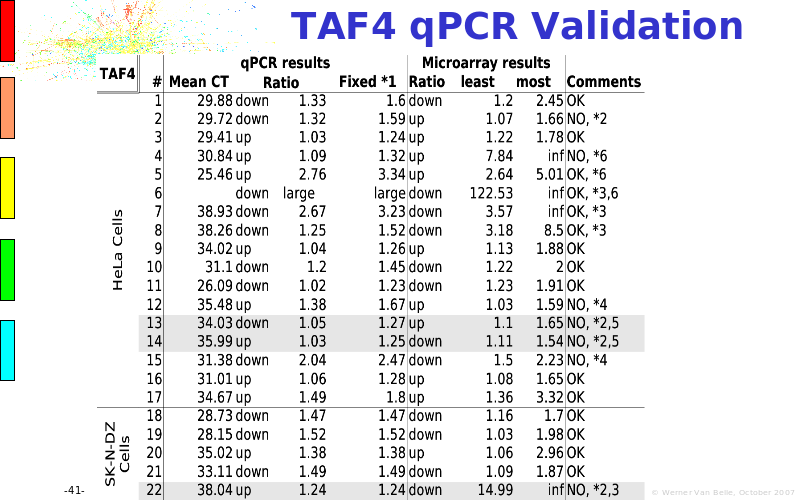
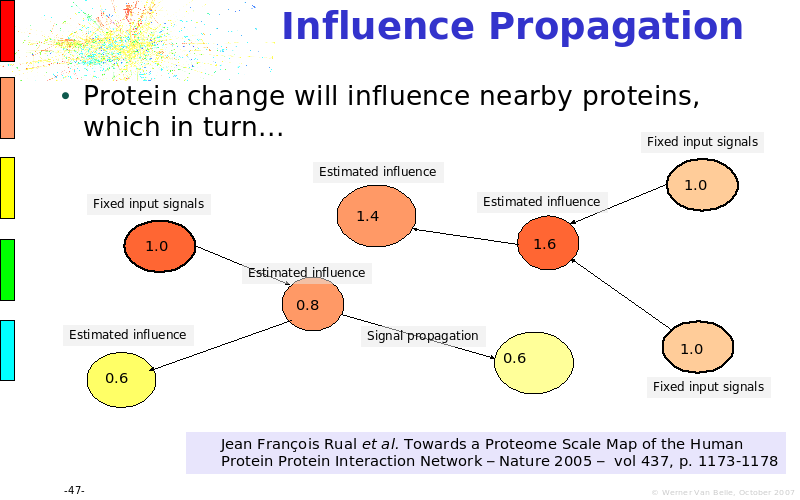
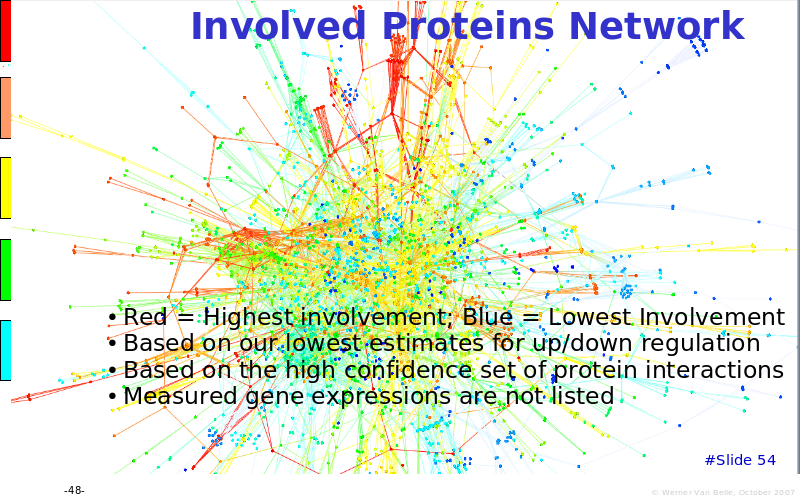
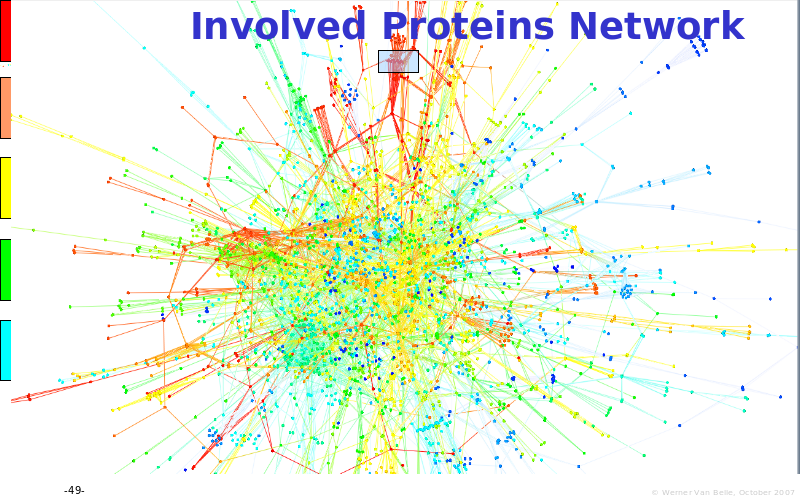
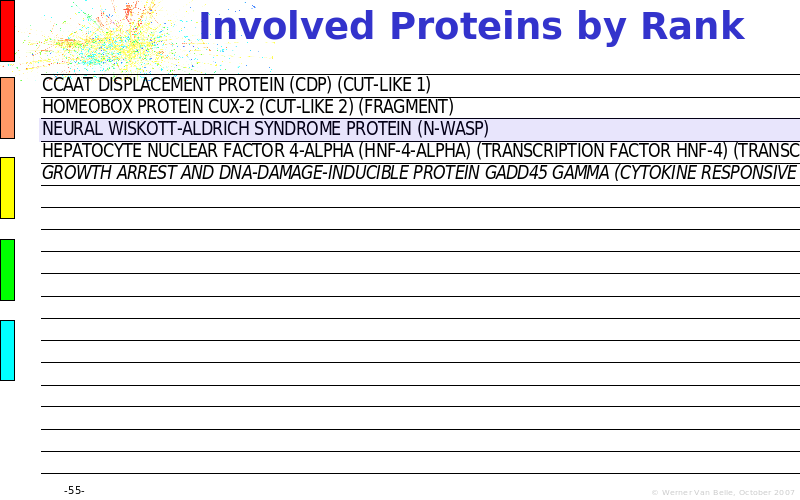
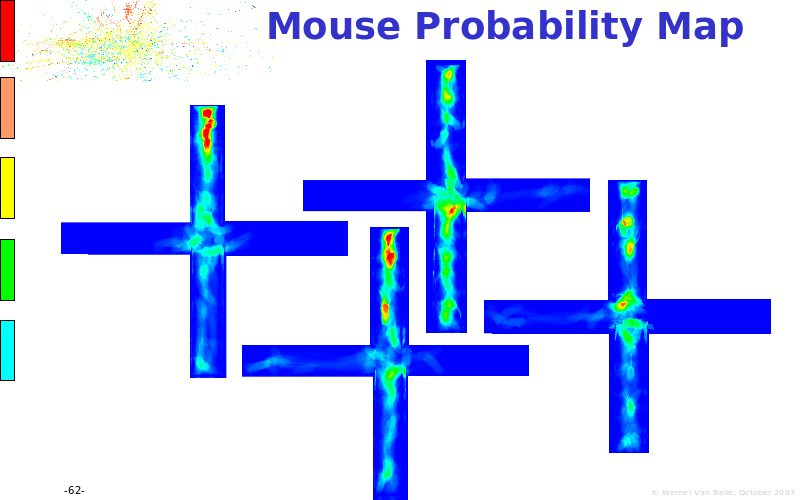
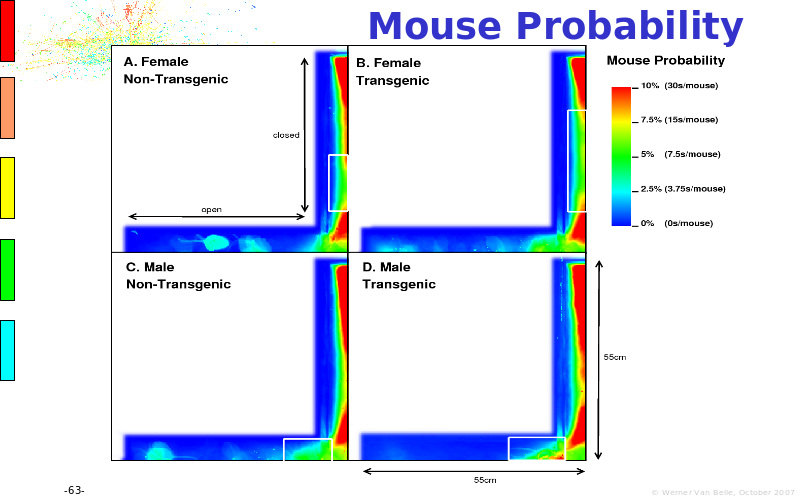
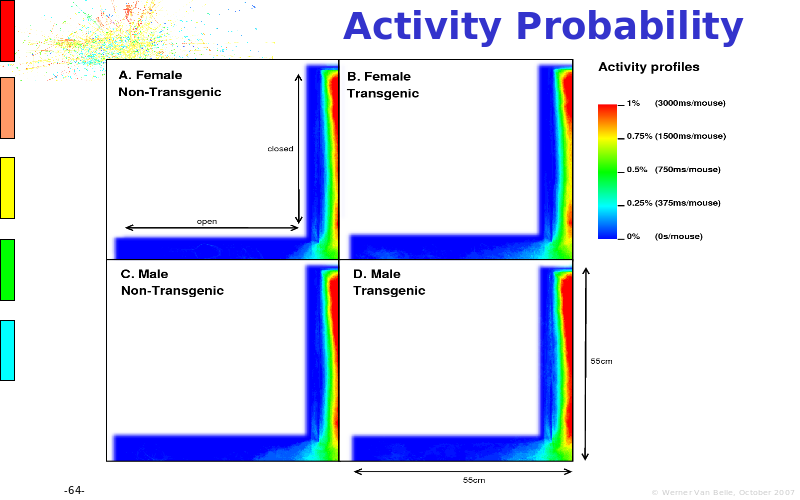
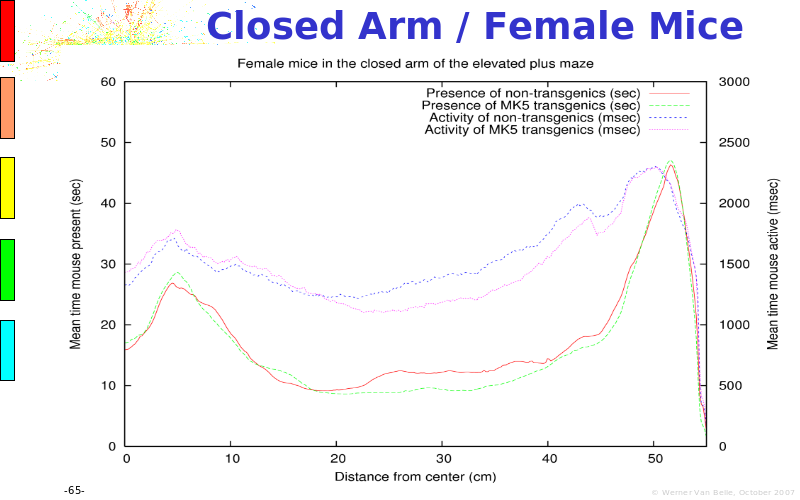
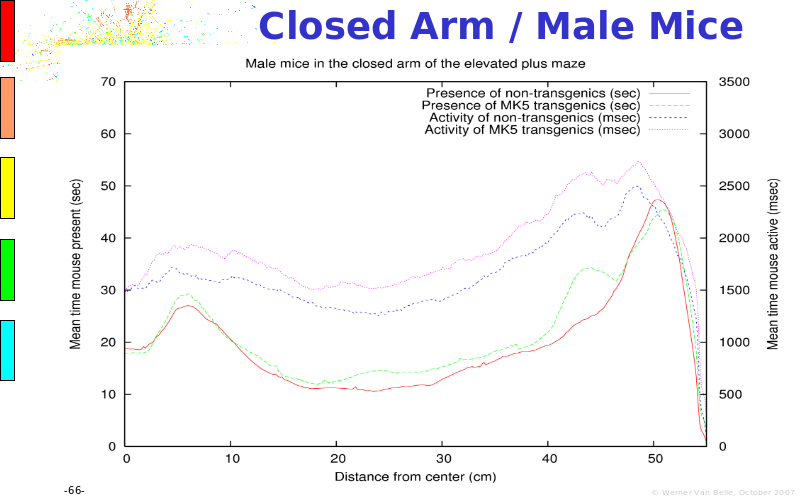
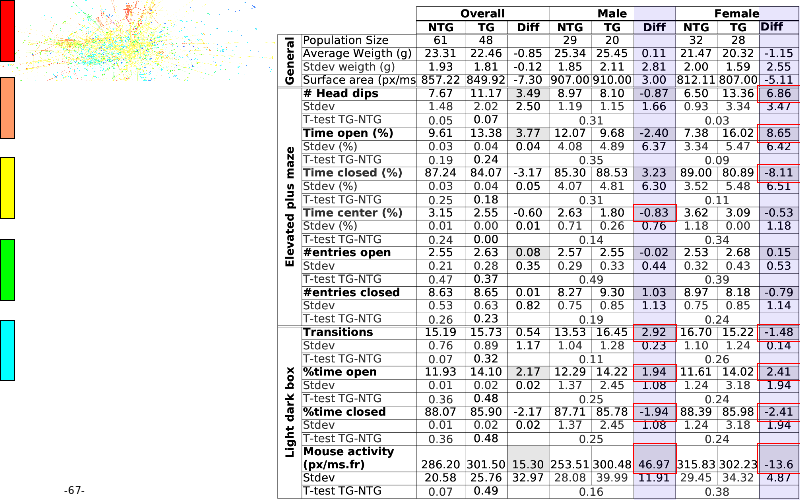
Abstract : The presentation consists of 3 parts. In the first part I present a novel method for the analysis of Microarrays. In brief, the method will, based on a number of replicates, create an intensity dependent error model. This model is then used to report confidence intervals on up/down regulation between different biological samples. The second part covers our use of these micro-array results by integrating them into a protein interaction map. A learning algorithm will then assess which impact the specific cell alteration potentially has on the proteome. The third part ellaborates on our functional validation of one of the predictions made by the reinforcement learner: MK5 influences anxiety. We validated this by means of transgenic mice on an elevated plus maze and in the light-dark box test
Keywords:
protein interaction maps, systems biology, micro array microarray measurement error
Reference:
Werner Van Belle; Microarrays, Protein Interaction Maps and Elevated Mazes; Presented at Animal Sciences Group, Wagingen Universiteit, Lelystad; The Netherlands; September 2007
Files: cropped.avi, diff.avi, active.avi, detected.avi, original.avi, mk5influence.tlp













































| http://werner.yellowcouch.org/ werner@yellowcouch.org |  |