| Home | Papers | Reports | Projects | Code Fragments | Dissertations | Presentations | Posters | Proposals | Lectures given | Course notes |
Biological Networks and Noise Propagation
Werner Van Belle1* - werner@yellowcouch.org, werner.van.belle@gmail.com
1- Yellowcouch;
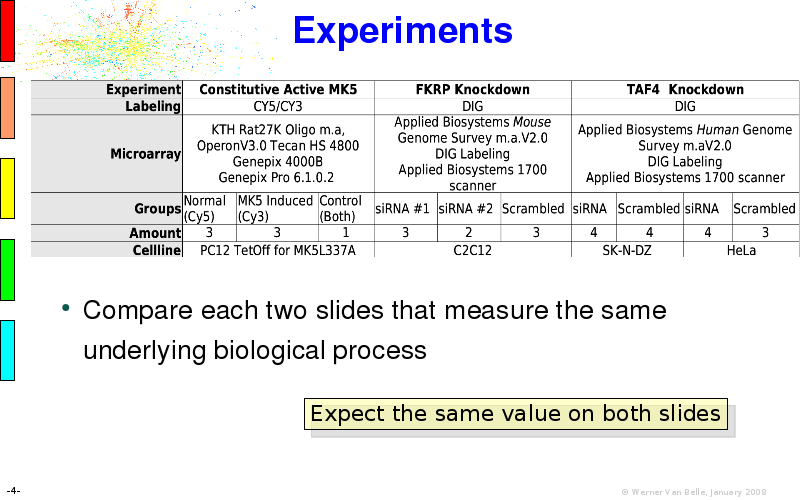
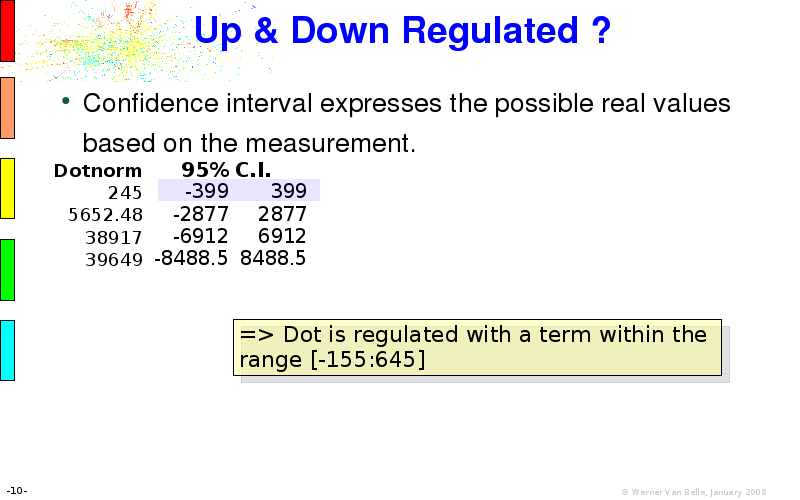
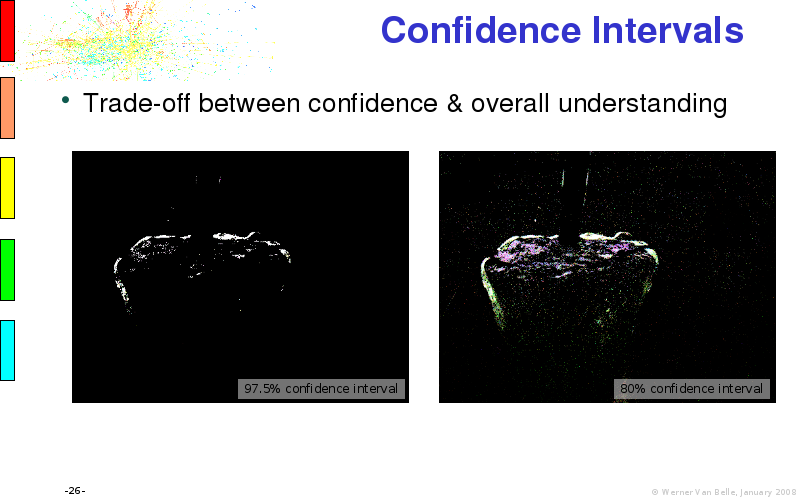
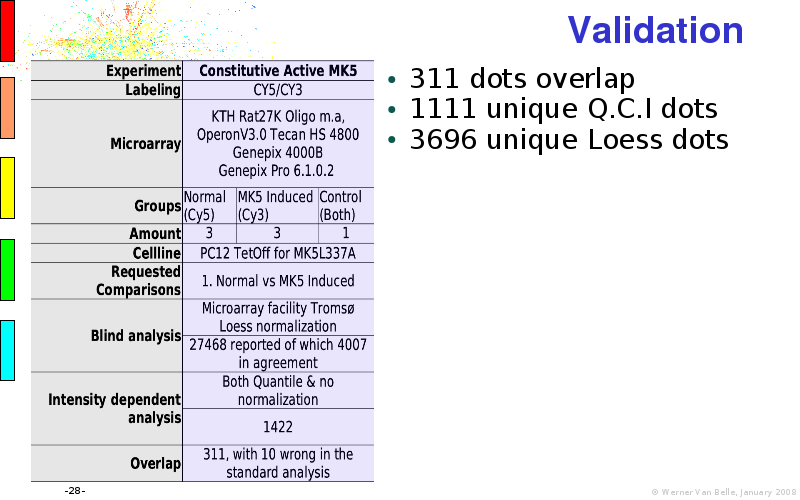
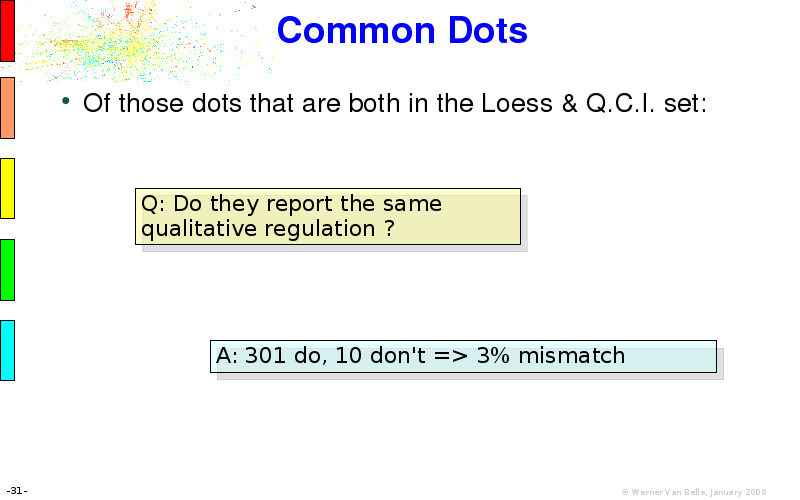
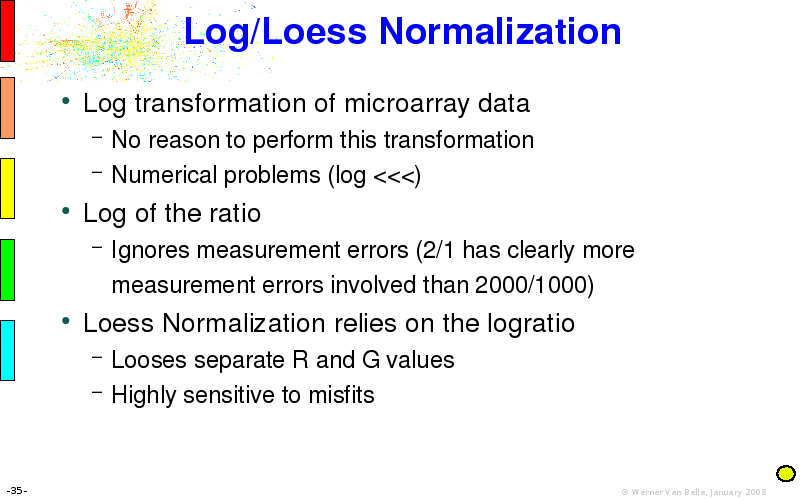
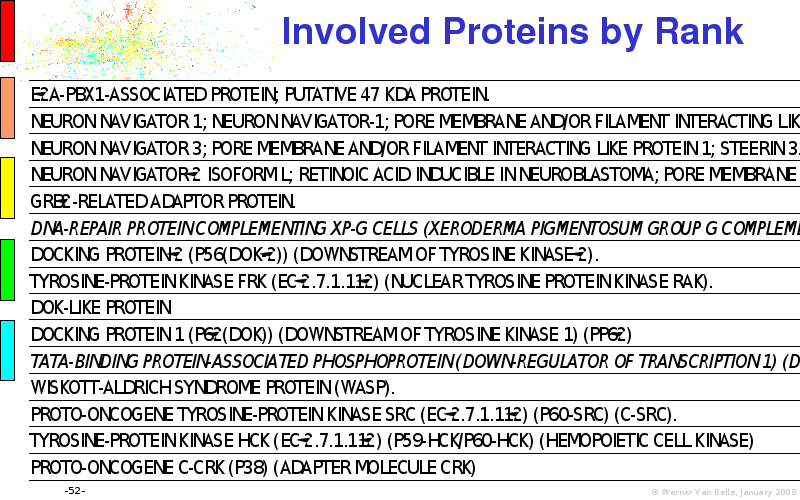
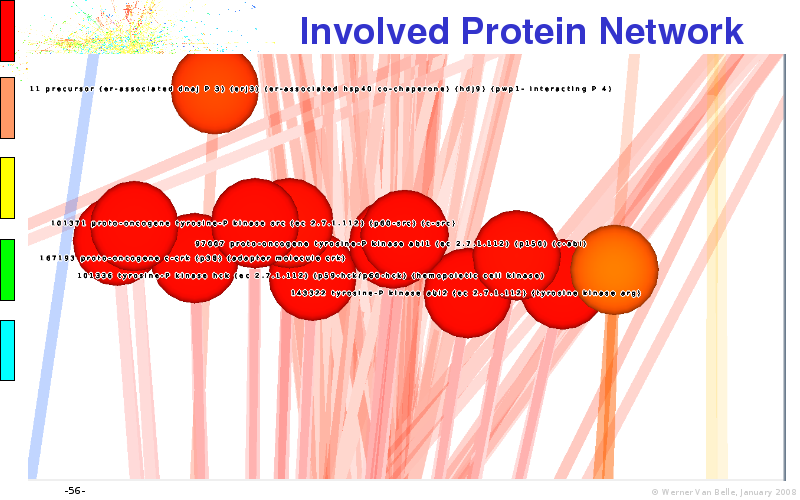
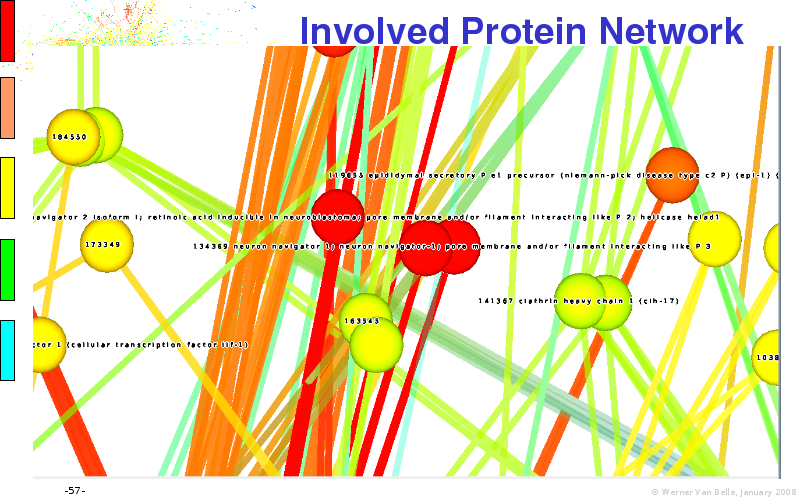
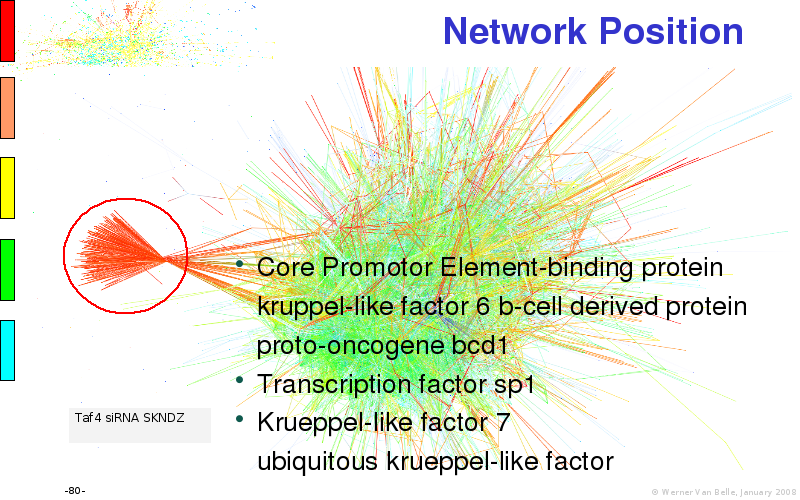
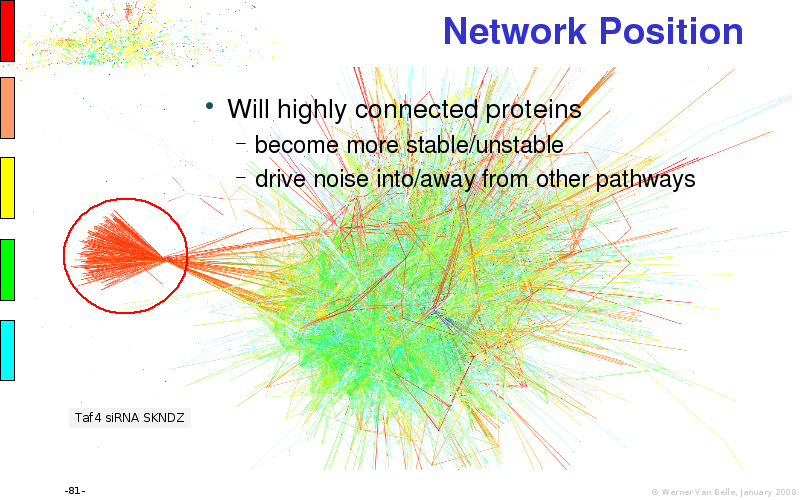
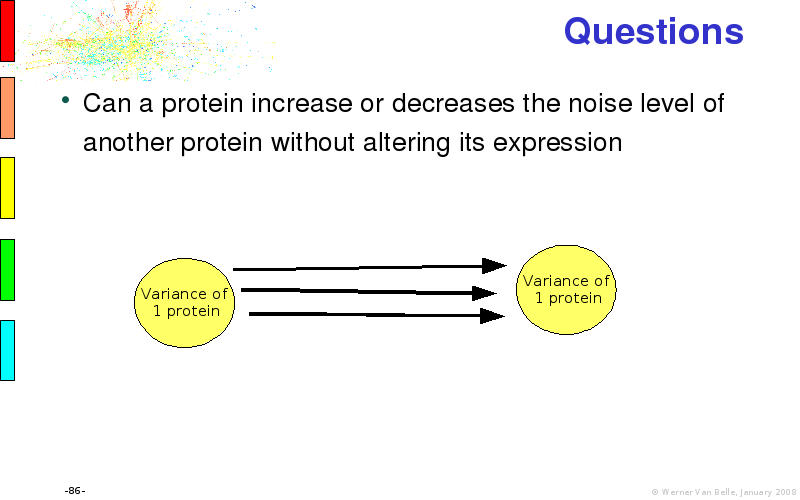
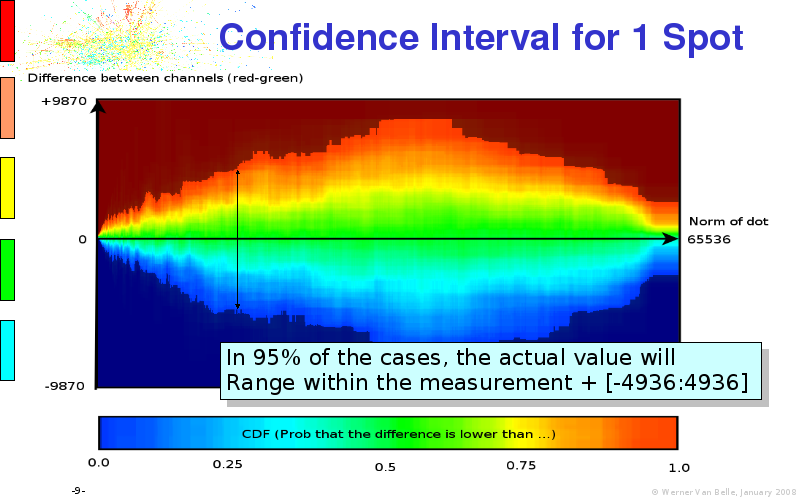
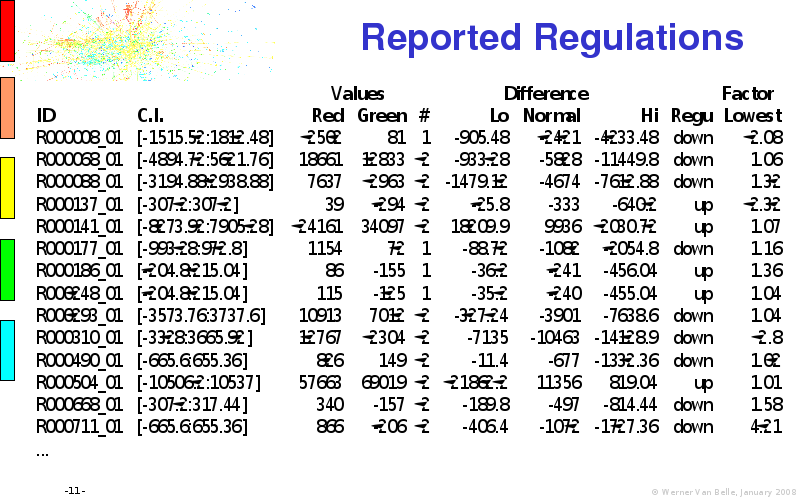
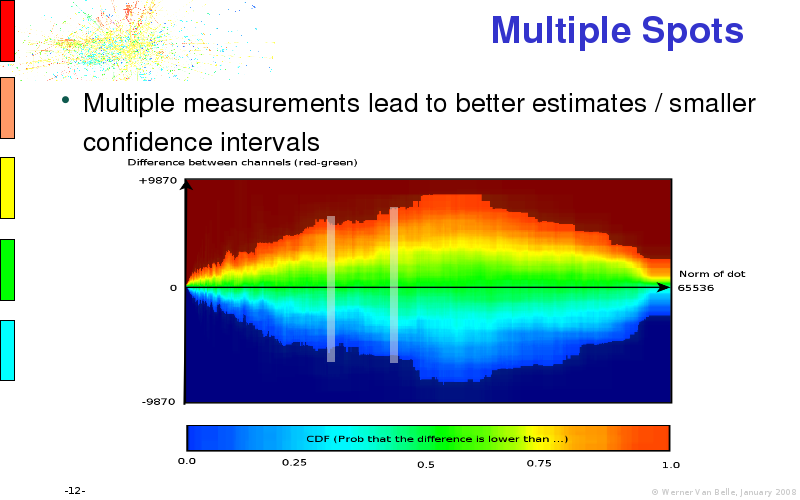
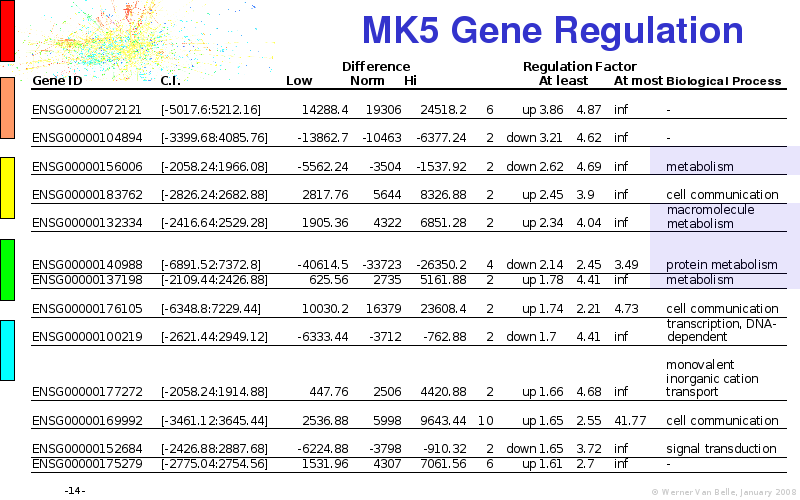
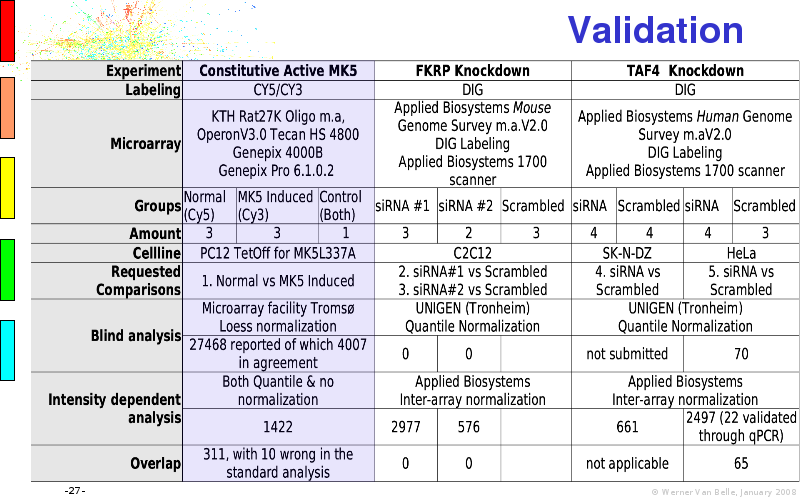
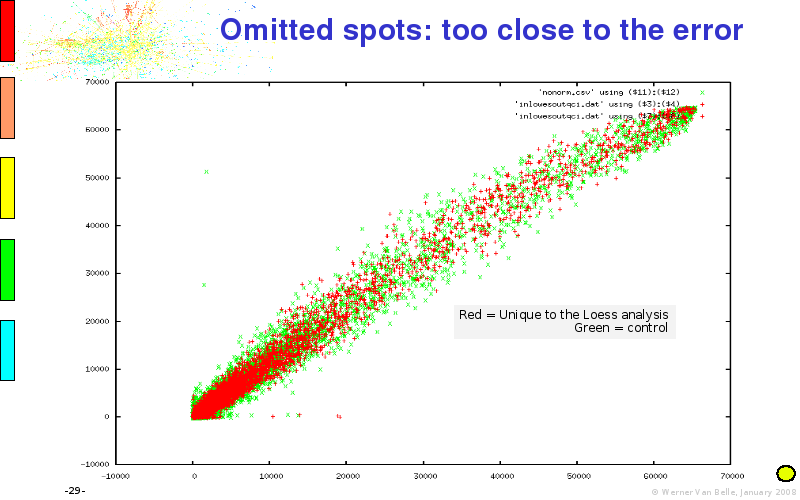
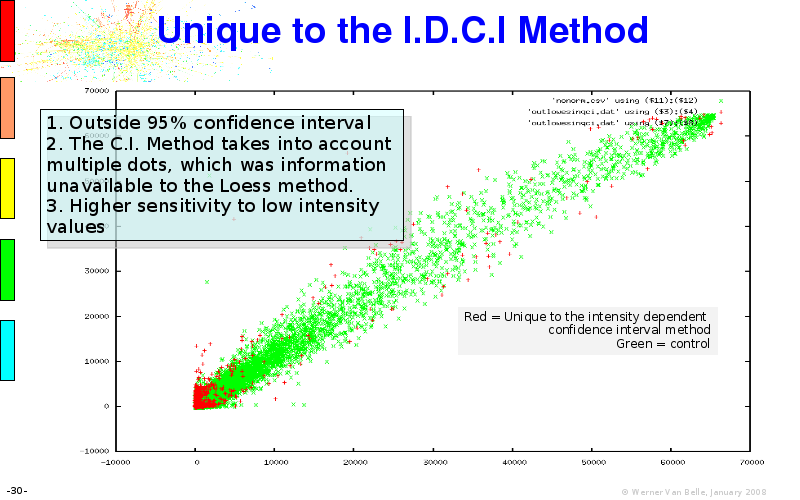
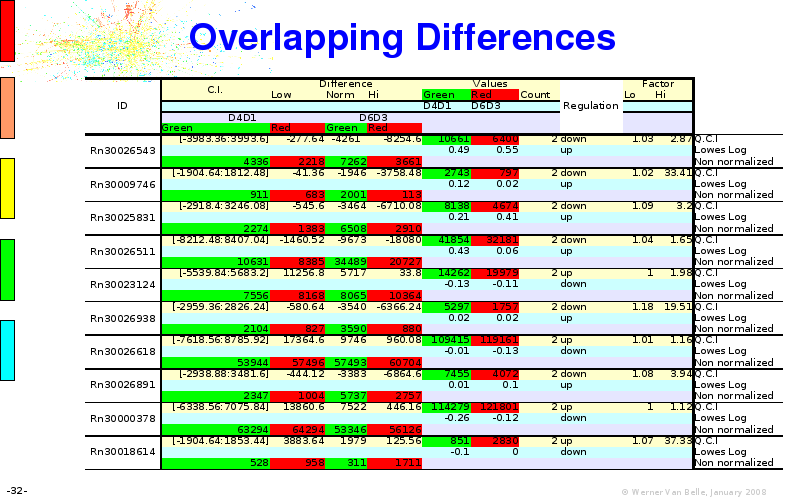
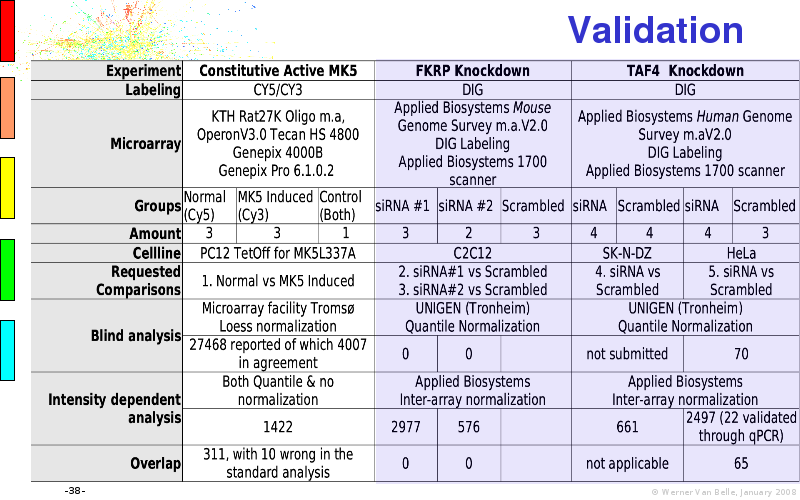
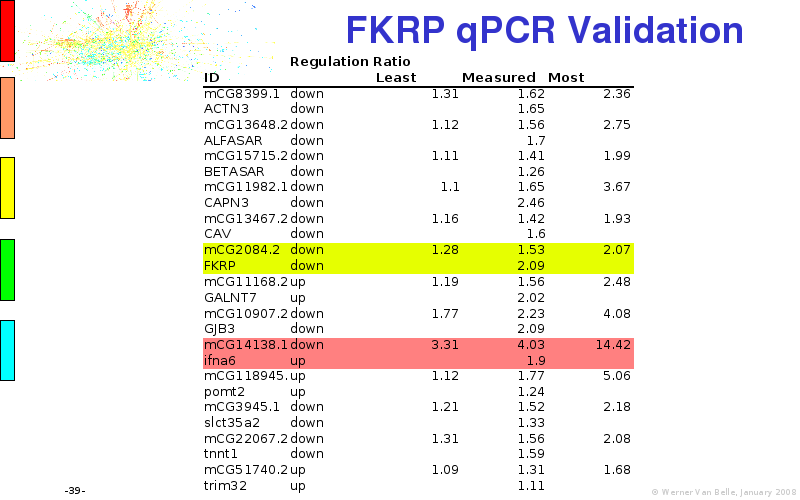
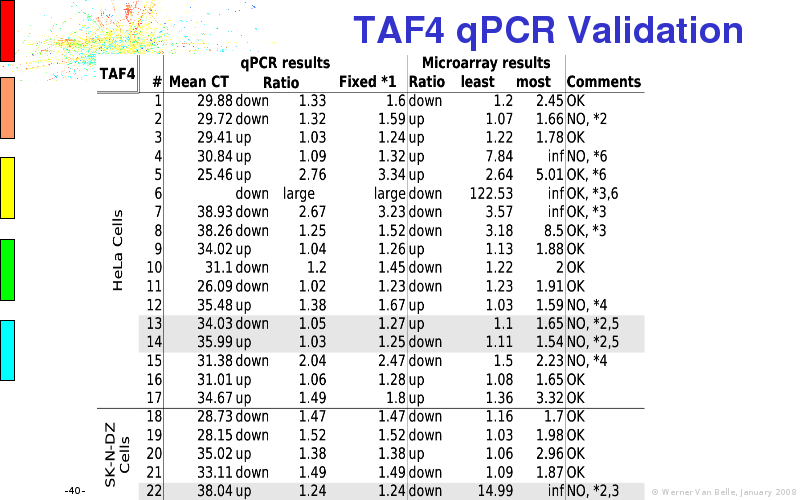
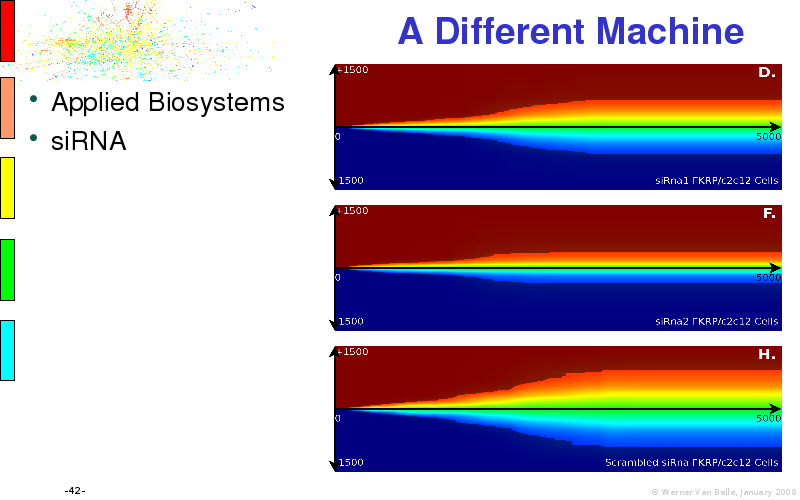
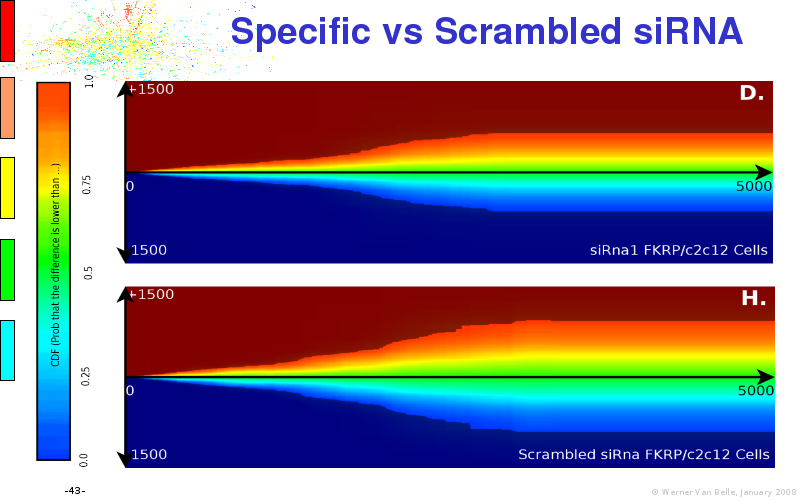
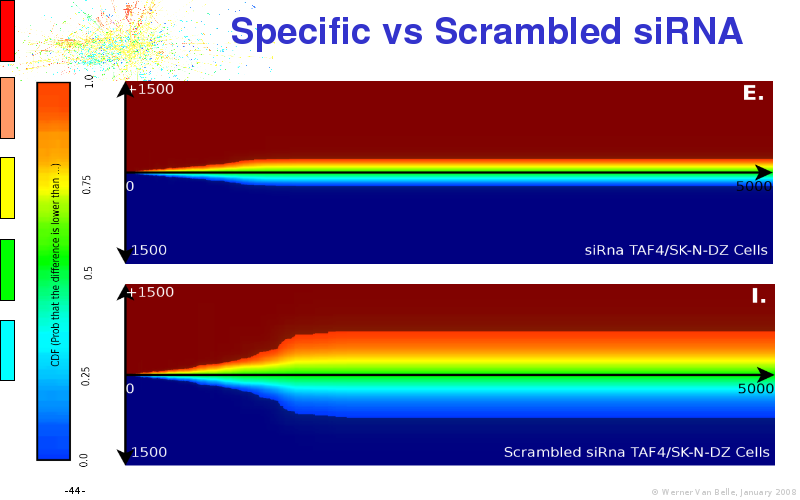
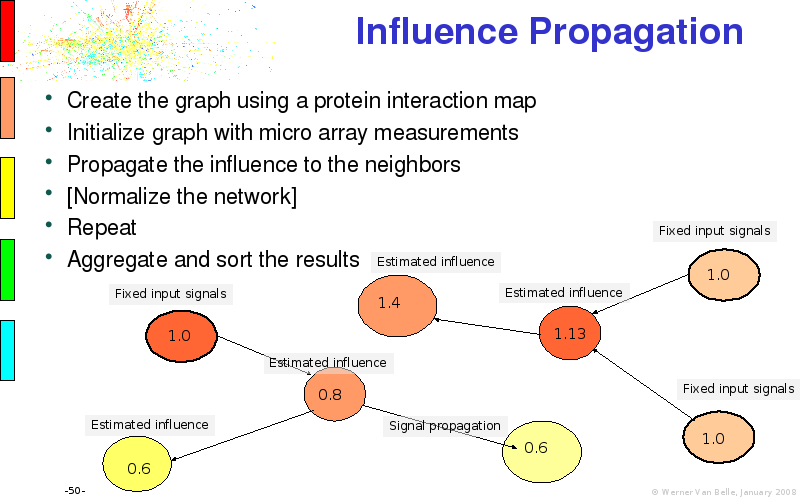
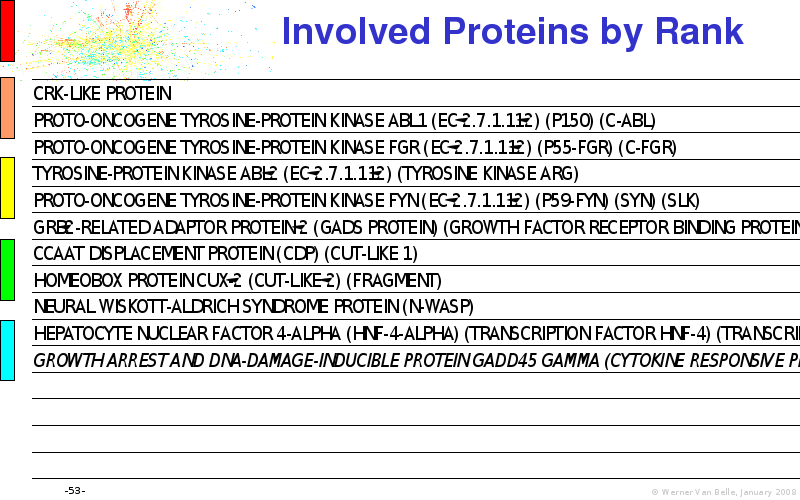
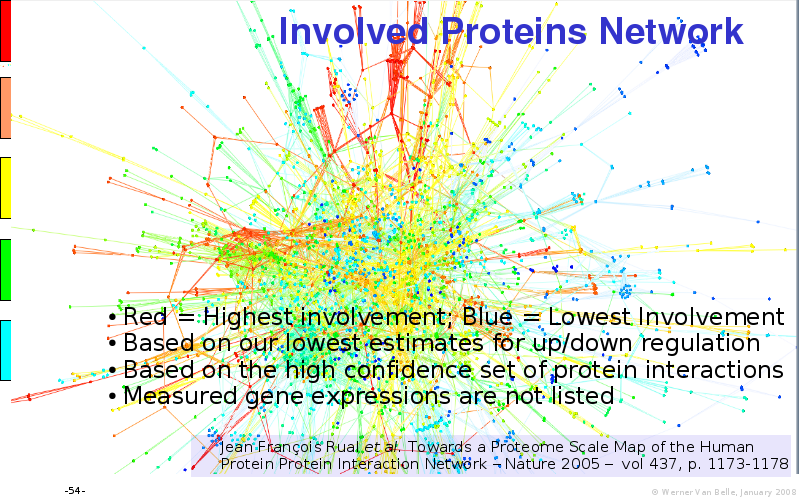
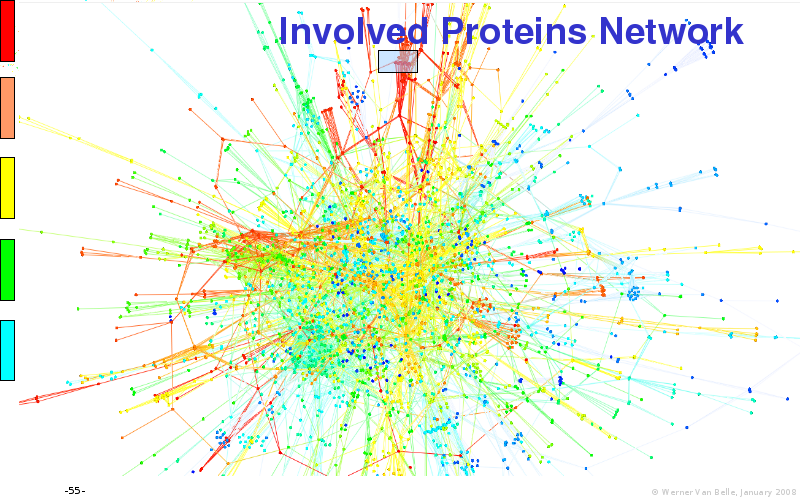
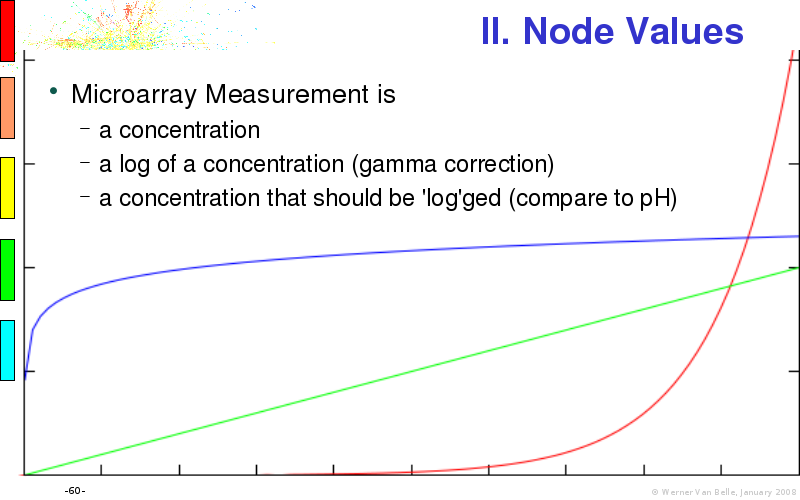
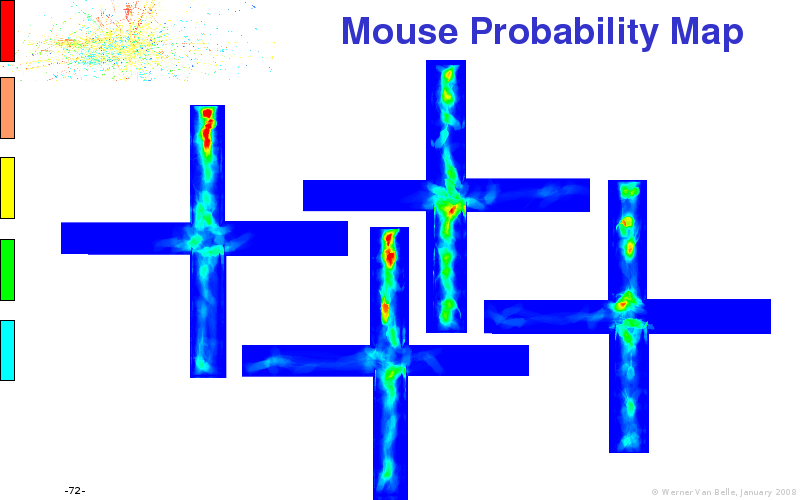
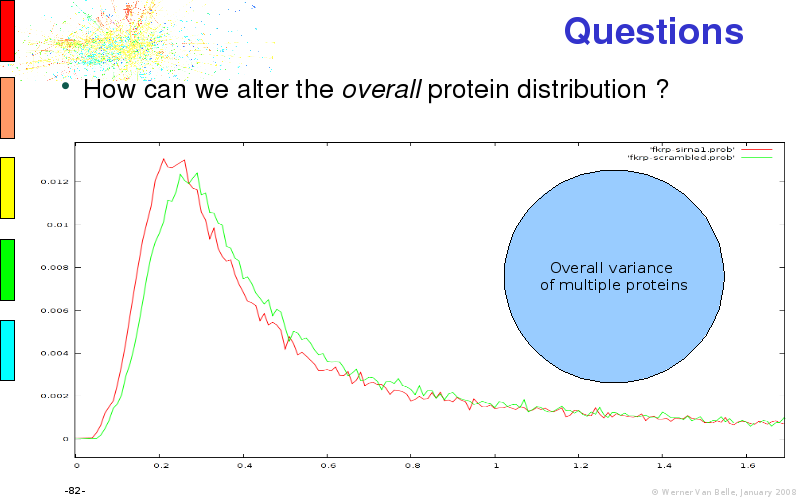
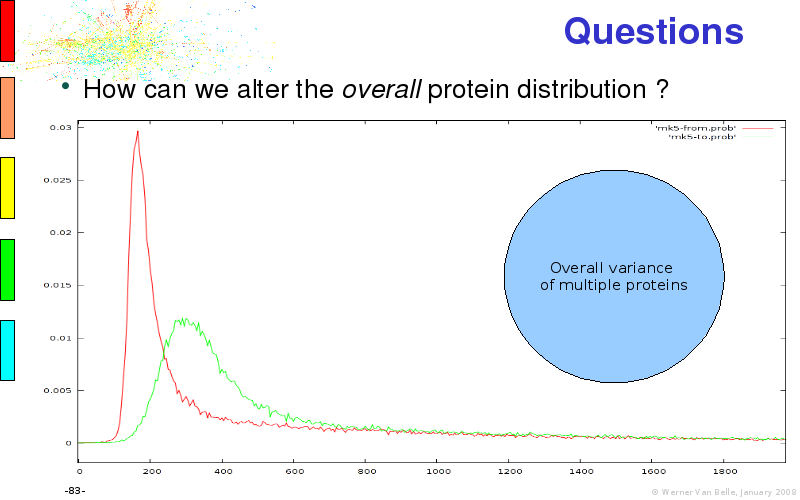
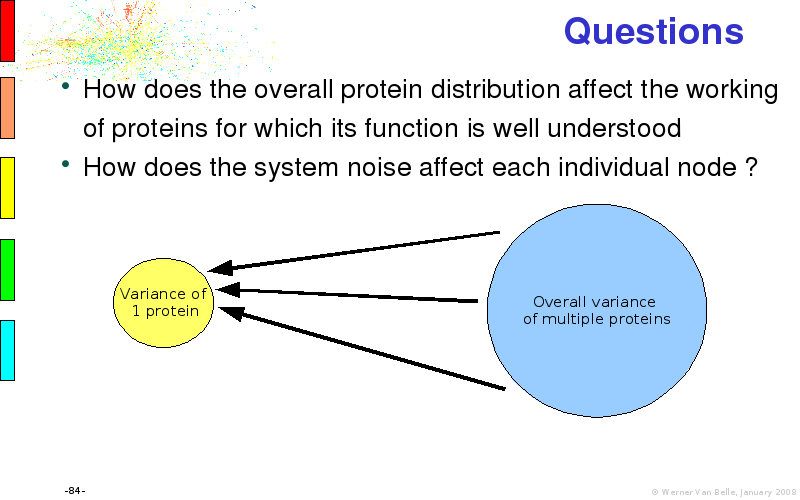
Abstract : I will present a) intensity dependent analysis of microarrays. This is an analysis technique that takes into account amplification errors and reports up/down regulations together with their 95% confidence intervals. I will explain the idea, the apparent improvement in analysis (x35 more genes reported) and the validation of this technique. The second topic focuses on b) assessing which protein are likely altered under specific gene knock-outs/knock-ins. We have been doing this by integrating a protein interaction map into micro array measurements and then propagating an influence value throughout that network. Although very rudimentary, this has given great results so far. I will give two example using MK5 and FKRP. I will wrap up the talk by focusing on normalization, measurement errors, biological variations and noise in genetic pathways.
Keywords:
noise propagation systems biology network protein interaction map integration microarray data visualization
Reference:
Werner Van Belle; Biological Networks and Noise Propagation; Presented at ETH Zurich in Basel; Switzerland; January 2008
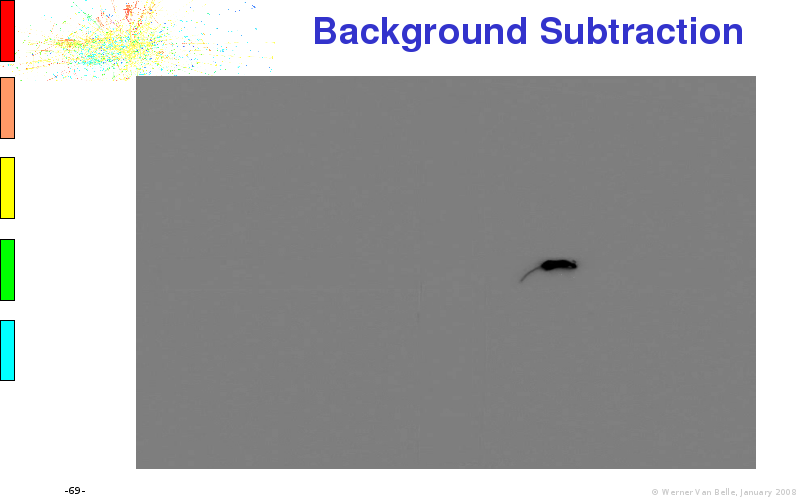
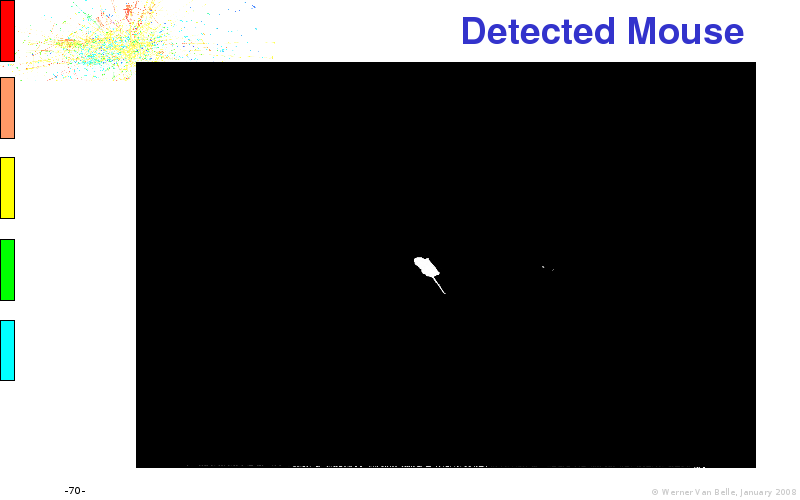
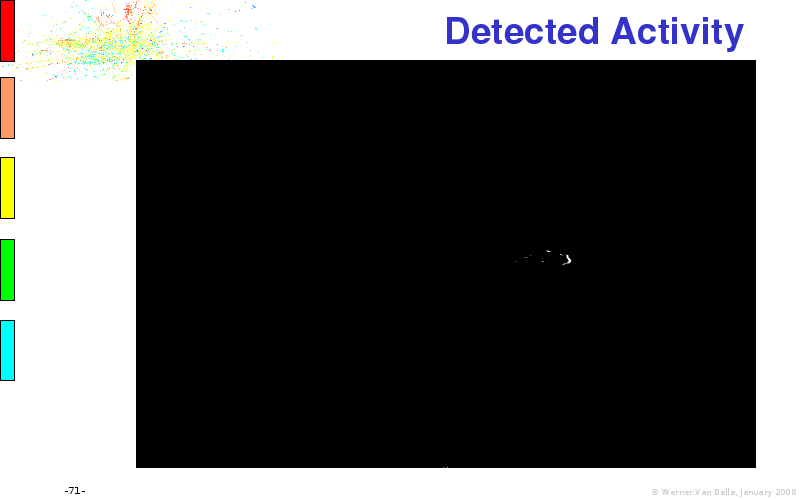
Files: original.avi, mk5influence.tlp, diff.avi, detected.avi, active.avi, cropped.avi
See also:
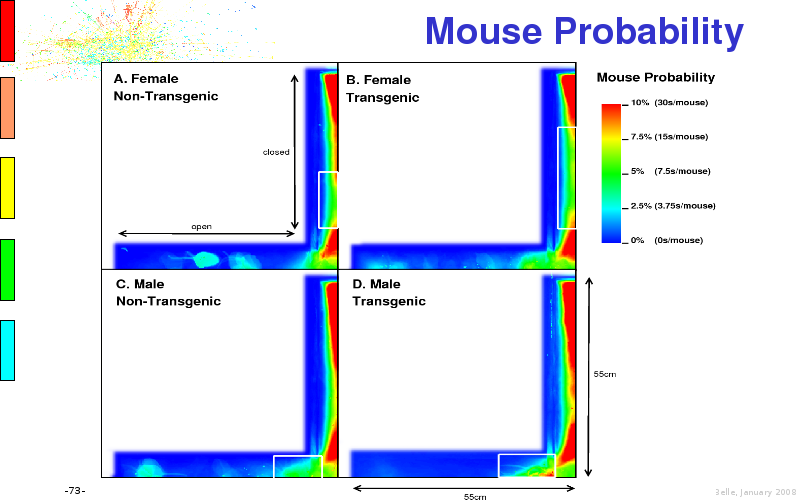
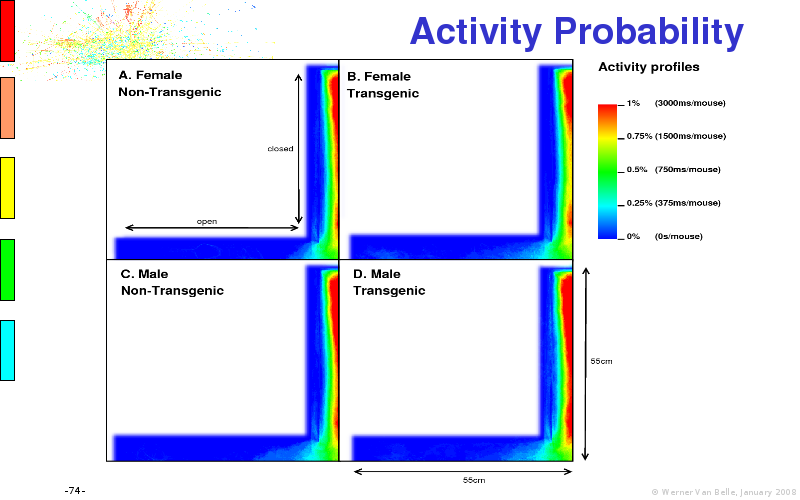
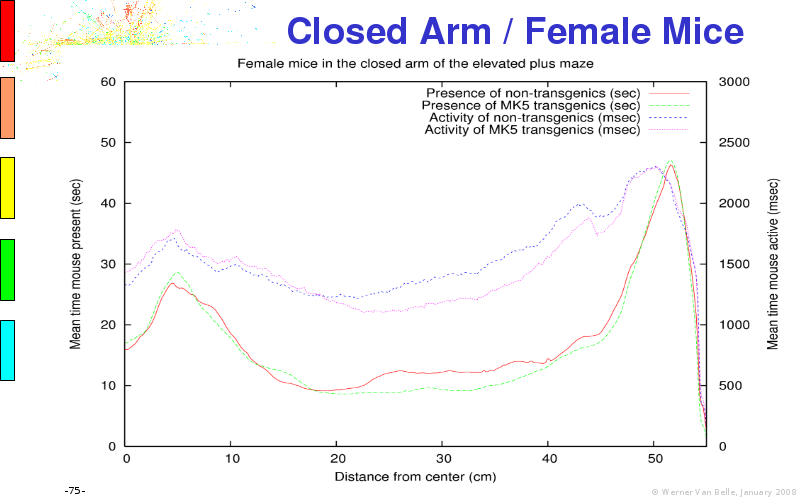
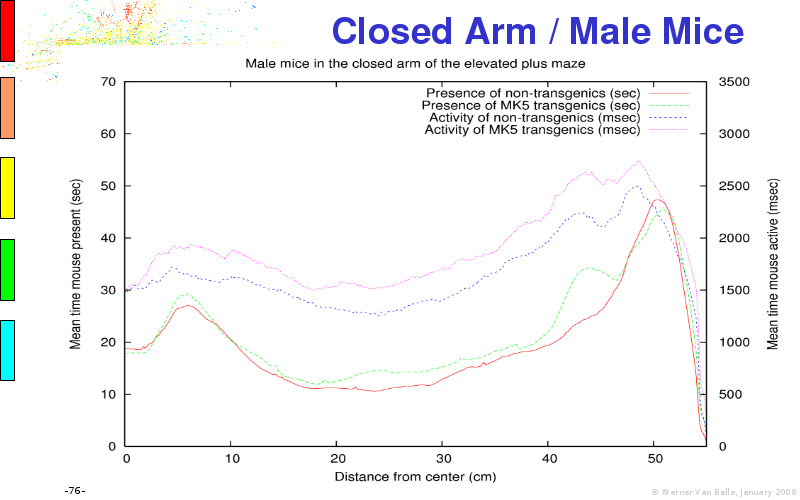
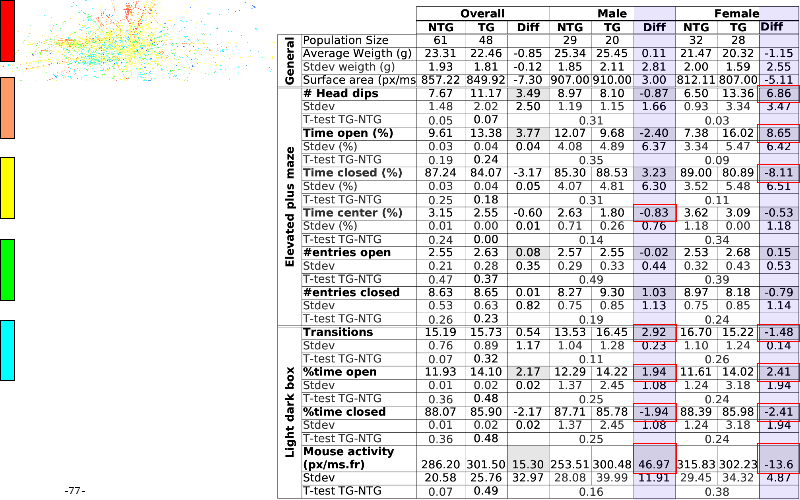
The article on anxious transgenic mice
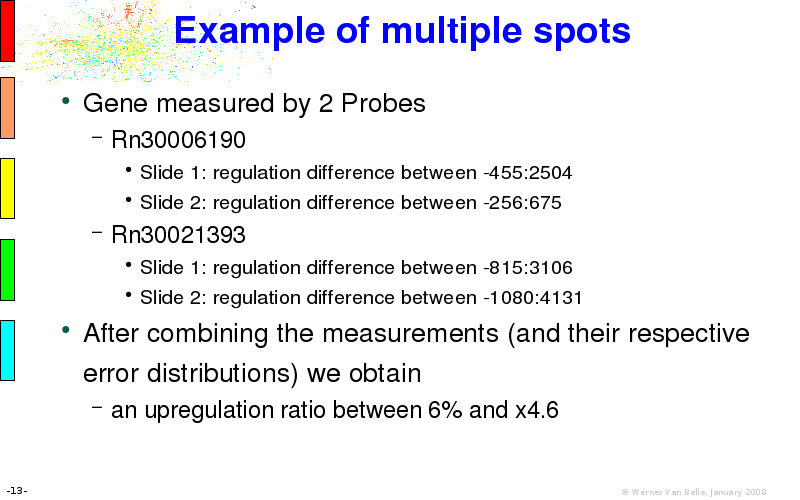
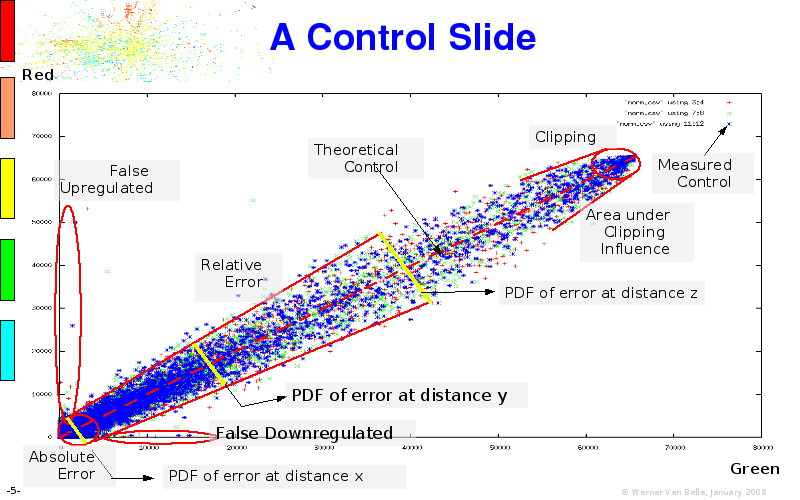
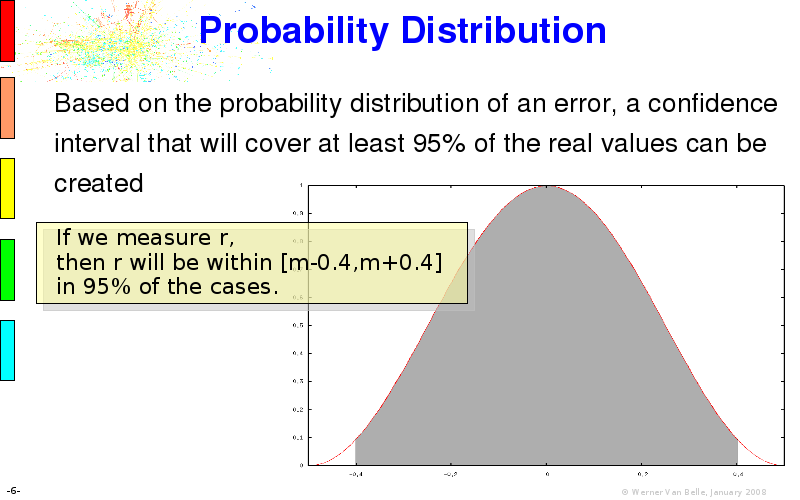
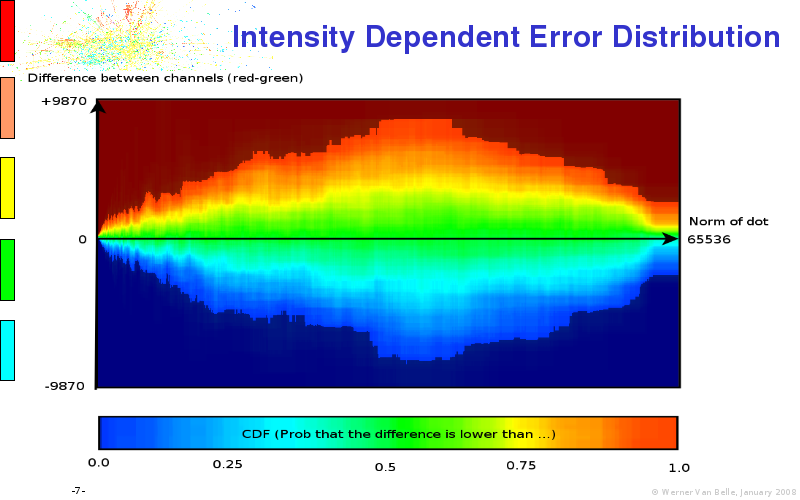
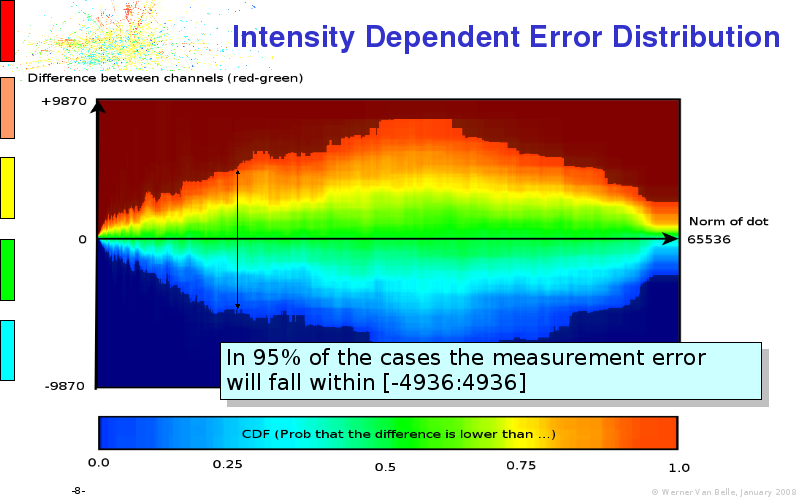
The article on intensity dependent confidence intervals




















































| http://werner.yellowcouch.org/ werner@yellowcouch.org |  |