| Home | Papers | Reports | Projects | Code Fragments | Dissertations | Presentations | Posters | Proposals | Lectures given | Course notes |
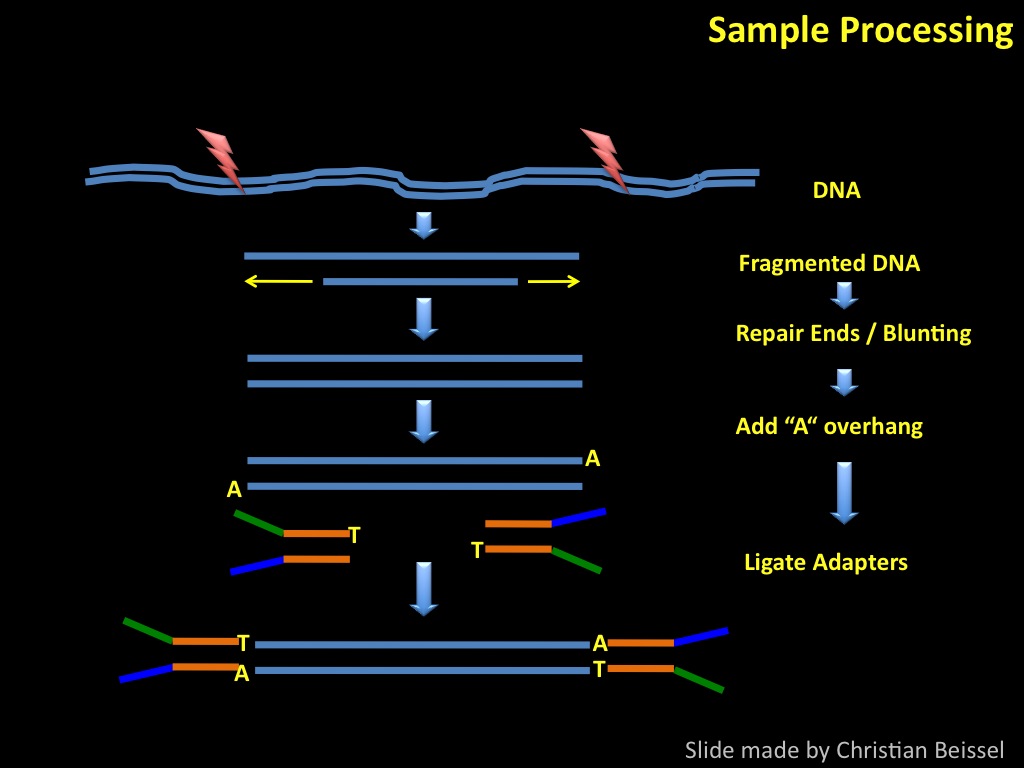
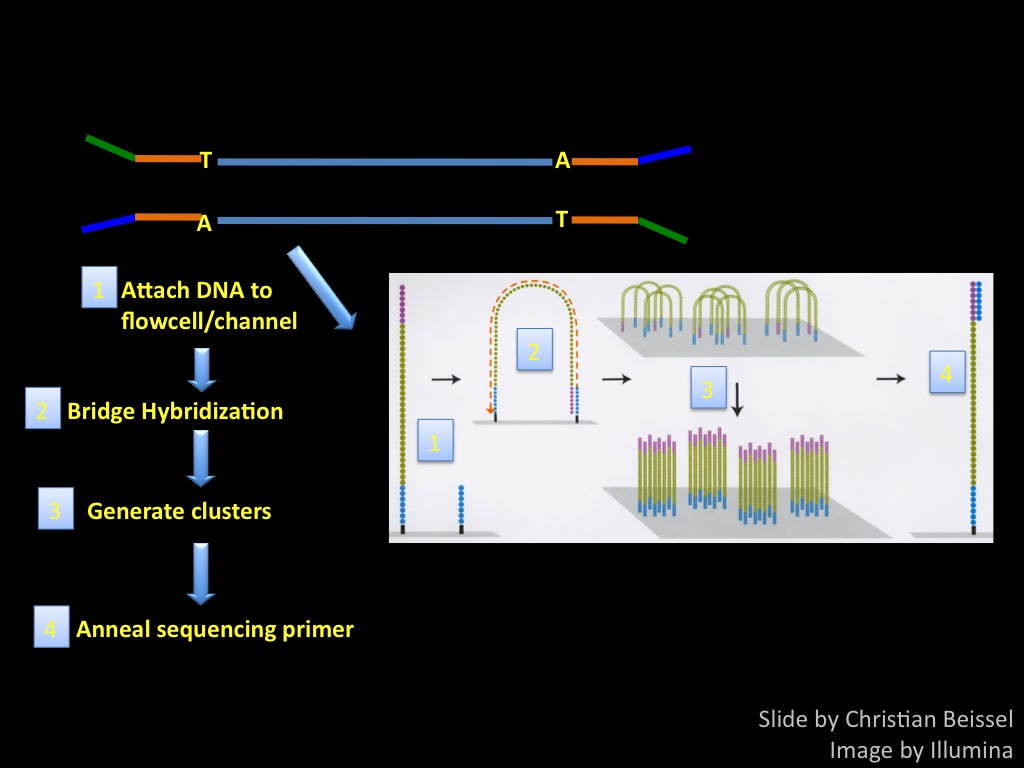
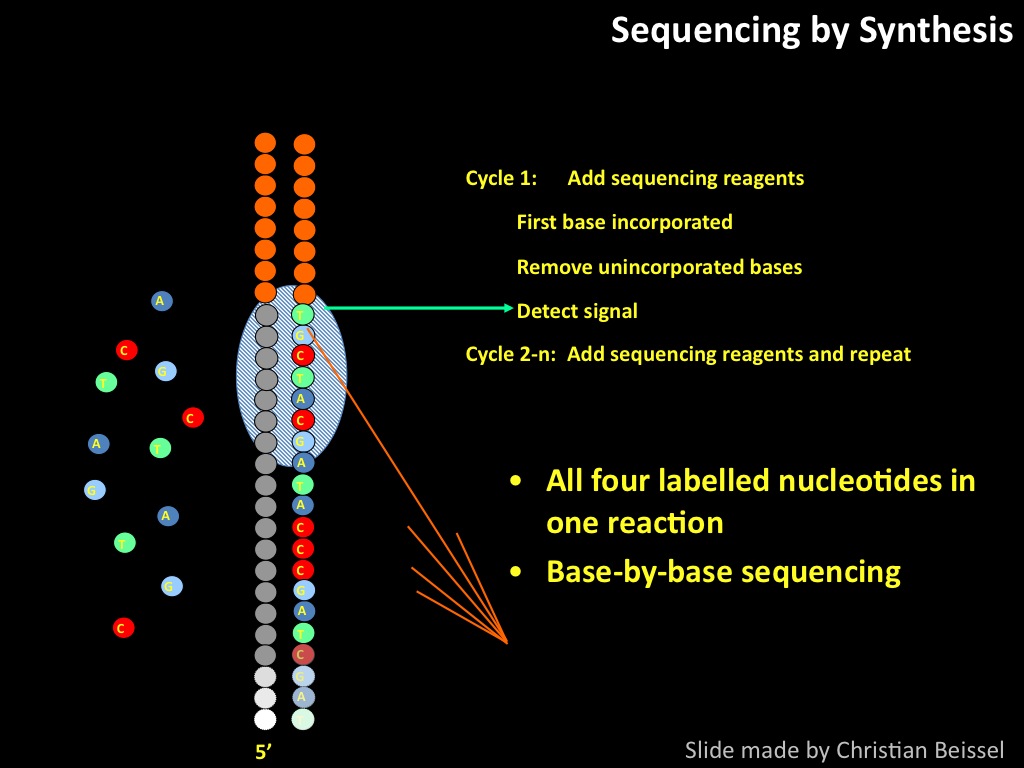
Sequencing by synthesis: from samples to data patterns.
Werner Van Belle1 - werner@yellowcouch.org, werner.van.belle@gmail.com
1- Deep Sequencing Unit
Department of Biosystems Science and Engineering
ETH Zurich; Mattenstrasse 24, Building 1058, Basel; Switzerland
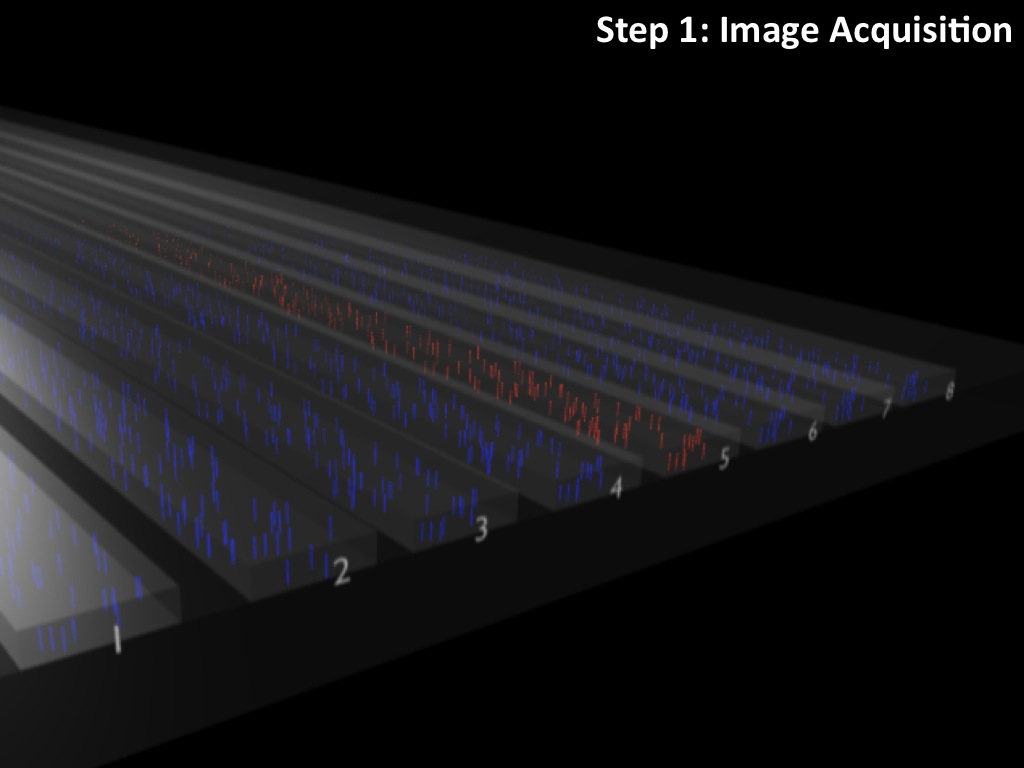
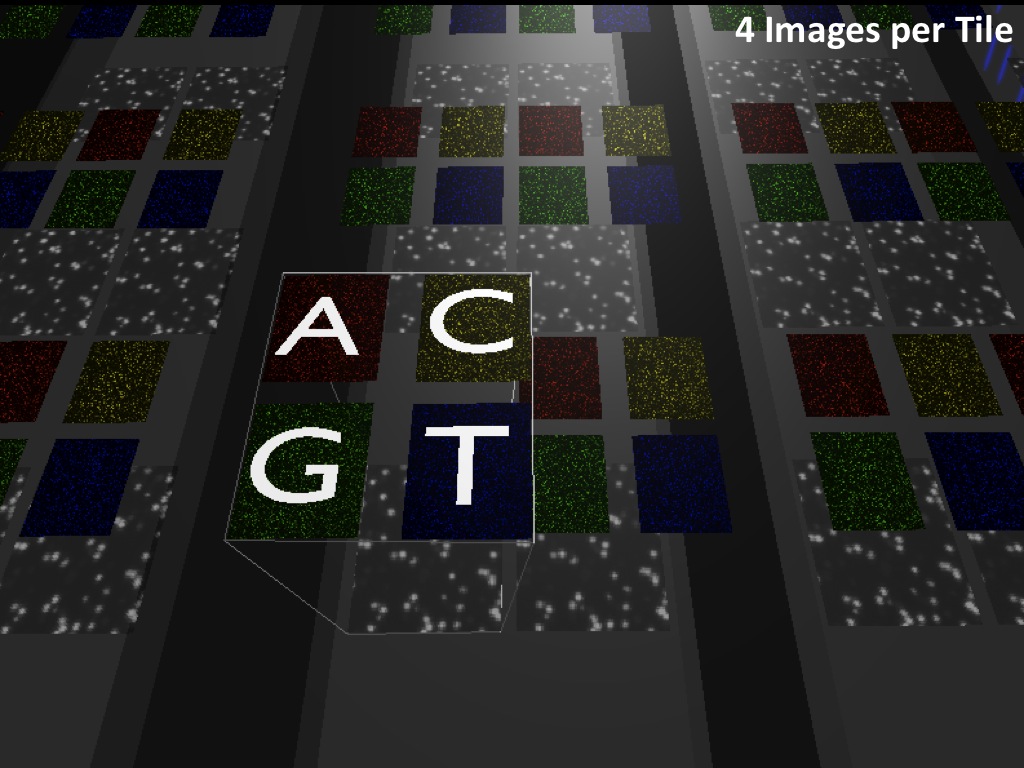
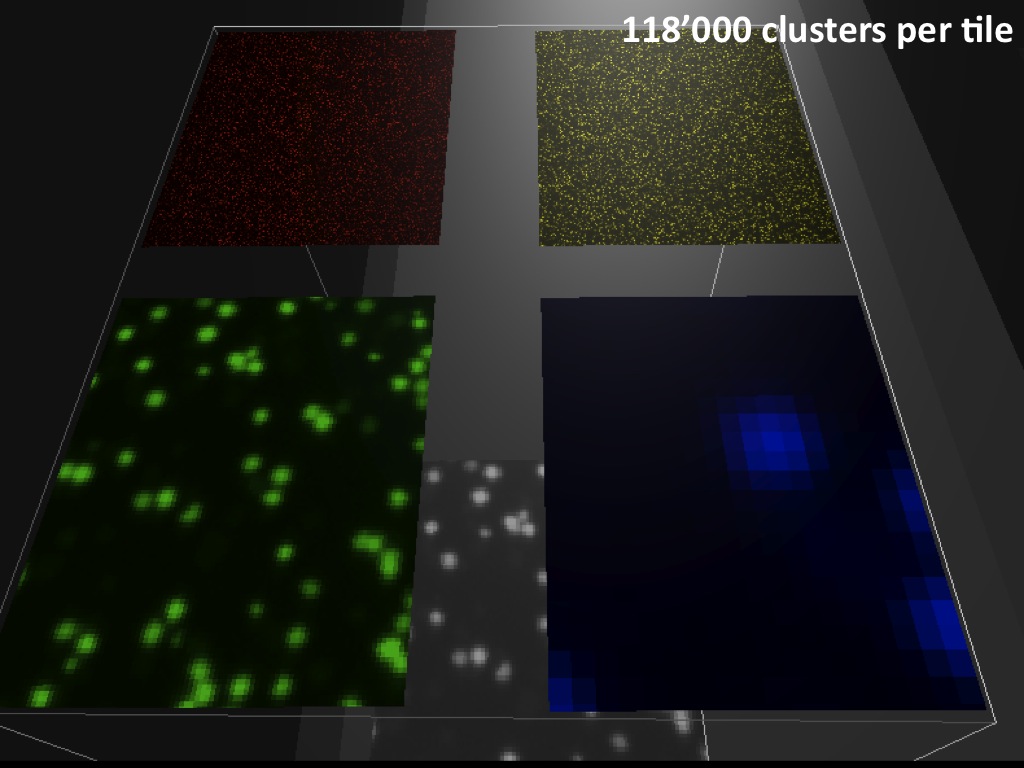
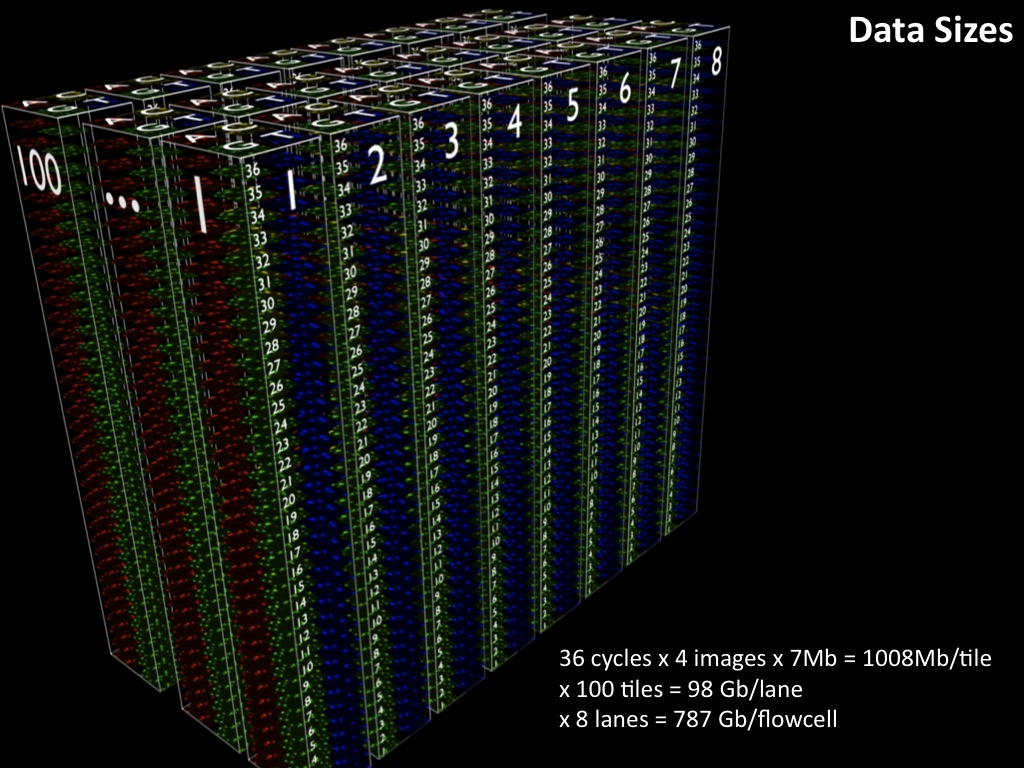
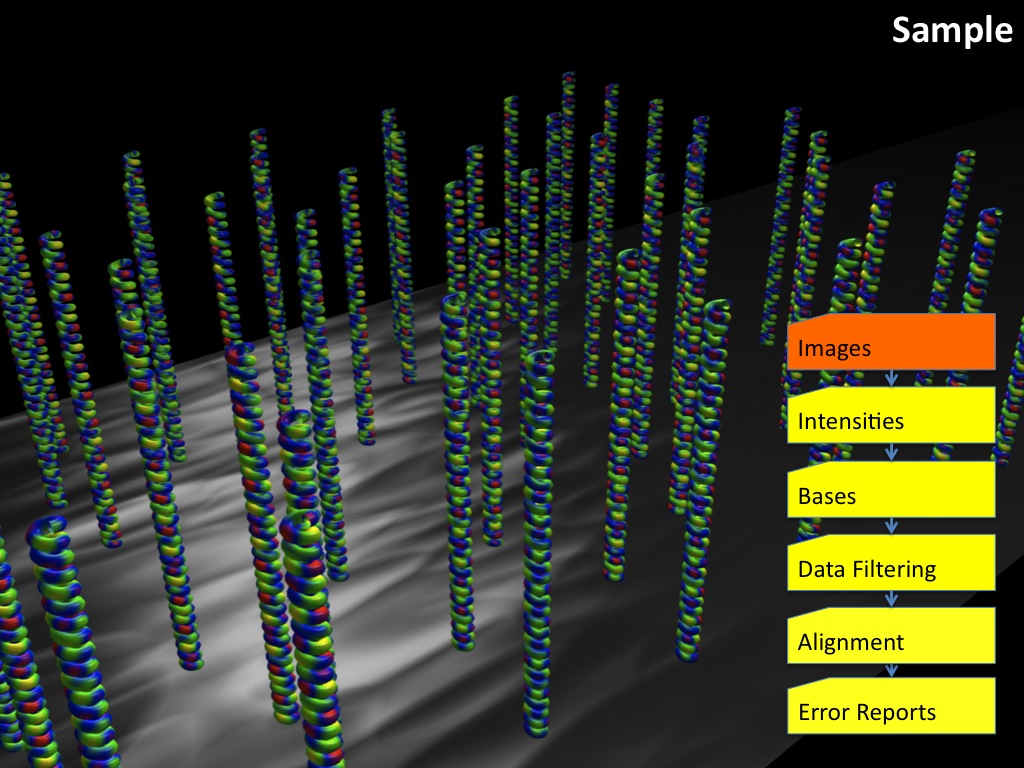
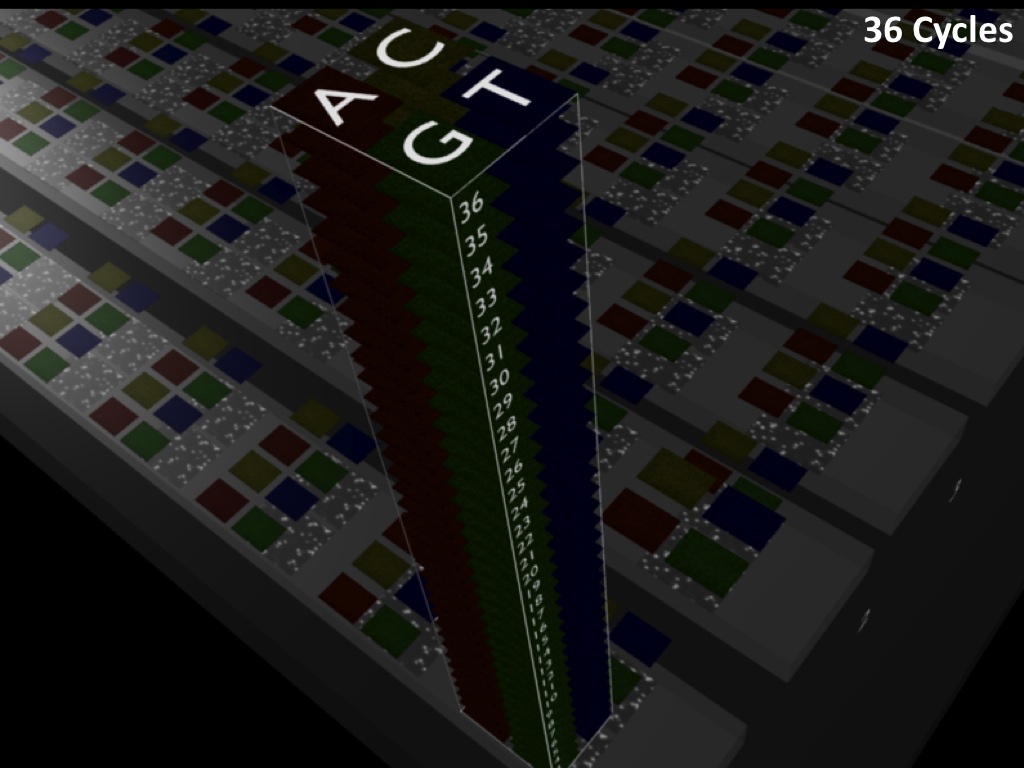
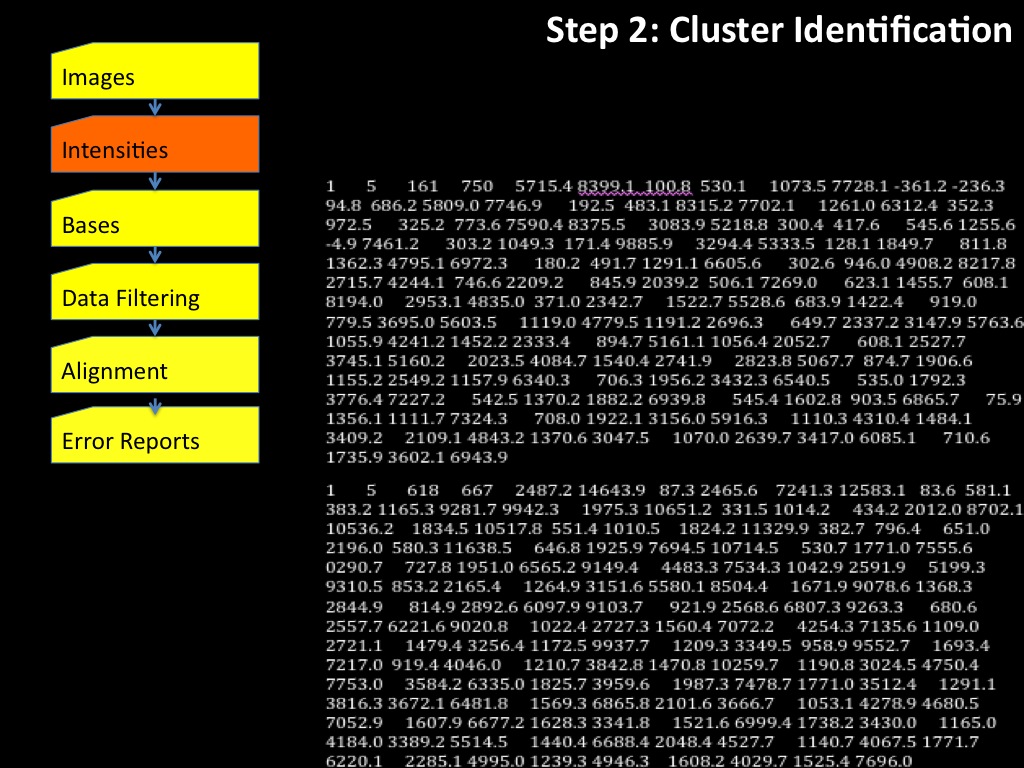
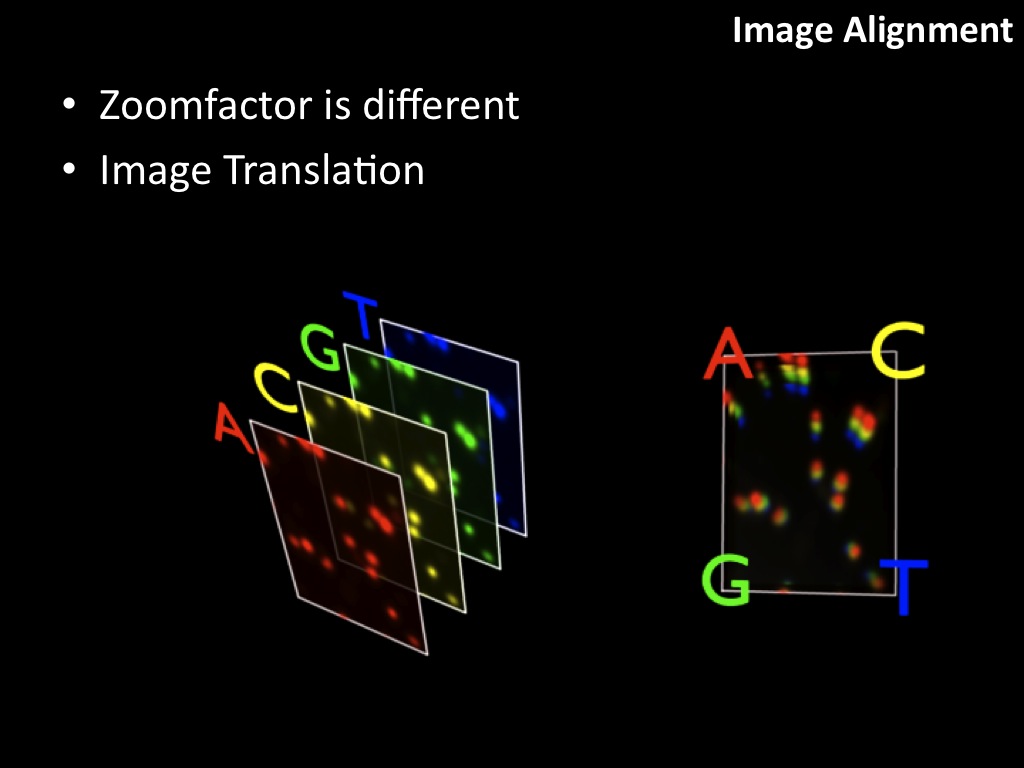
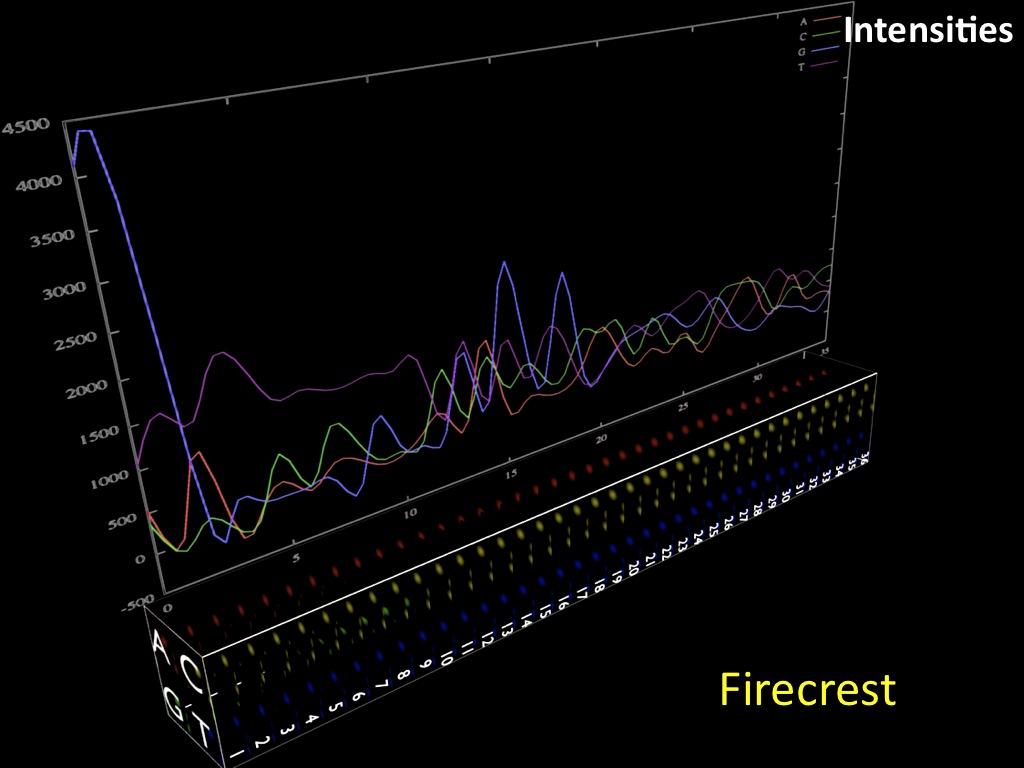
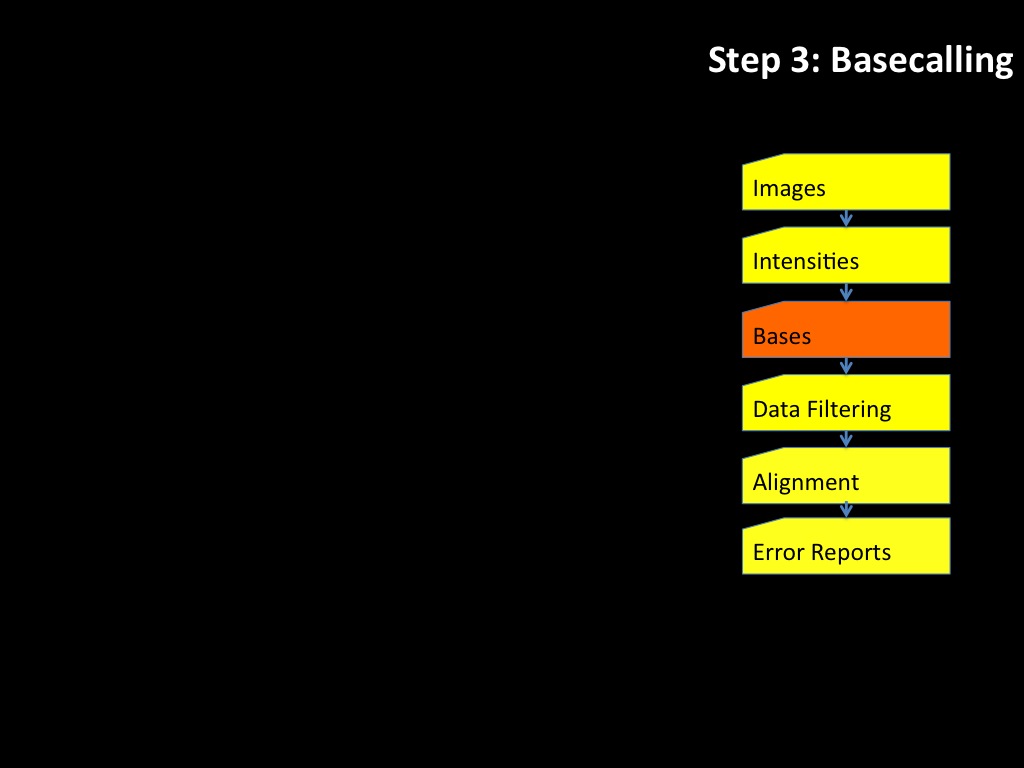
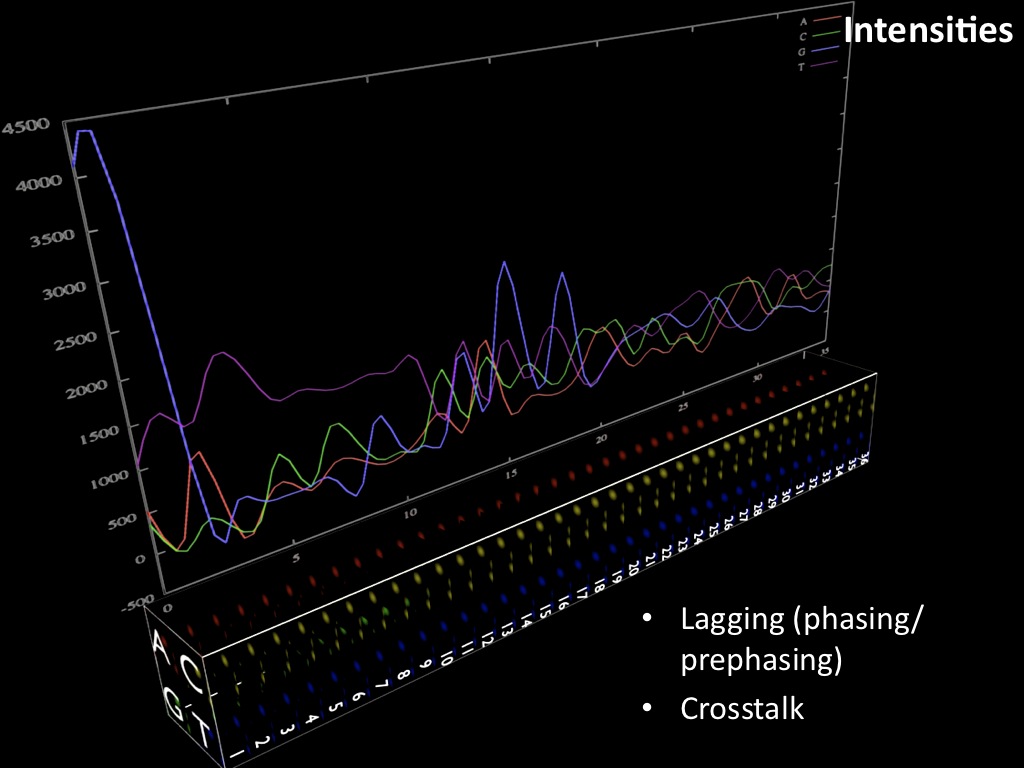
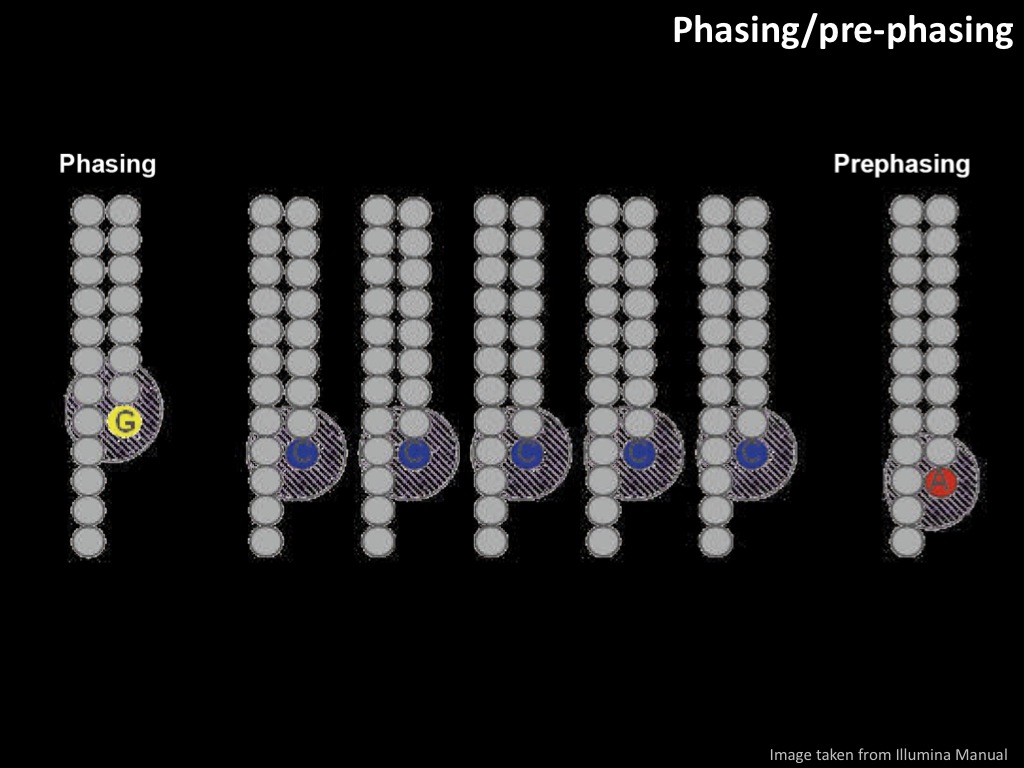
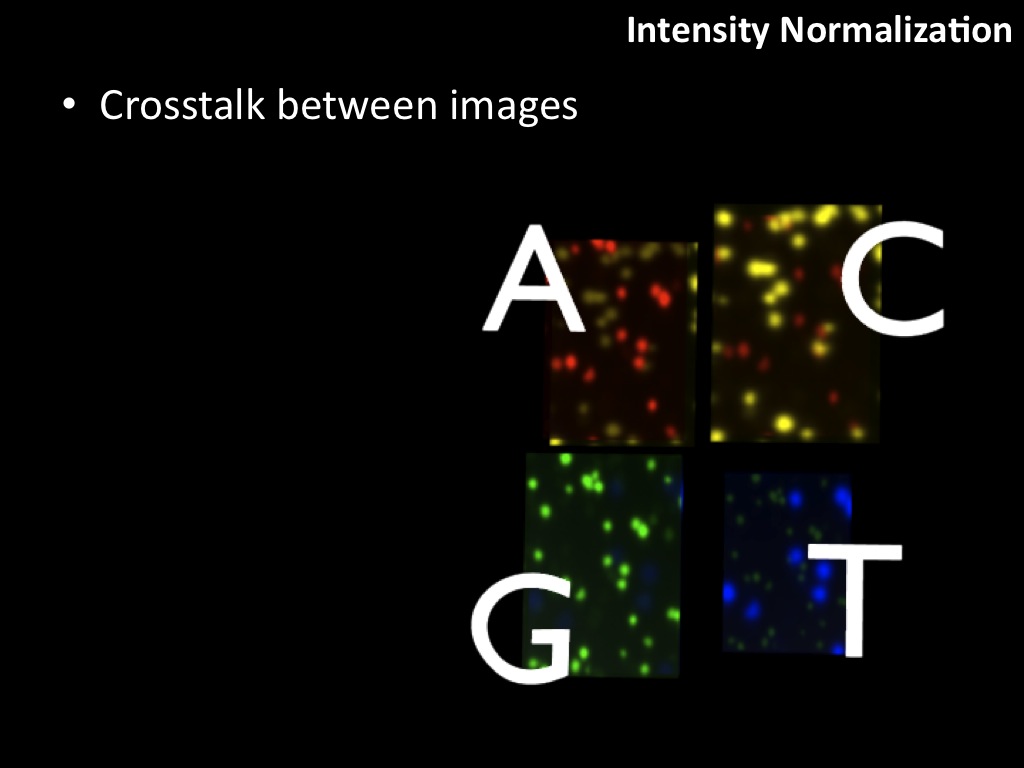
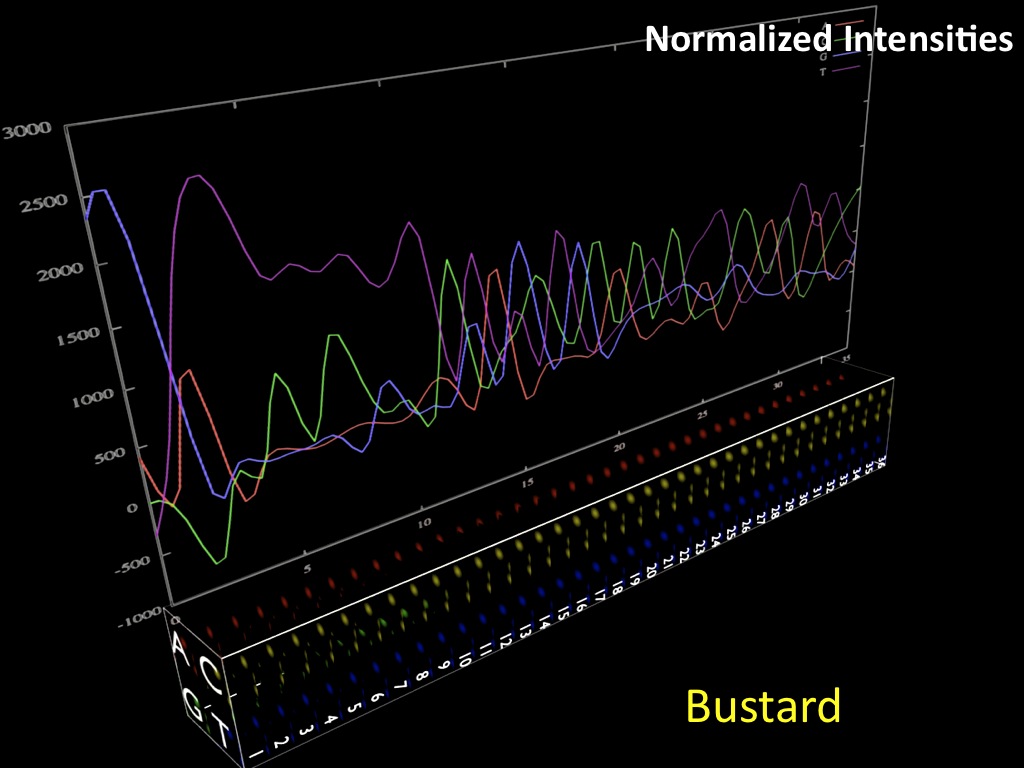
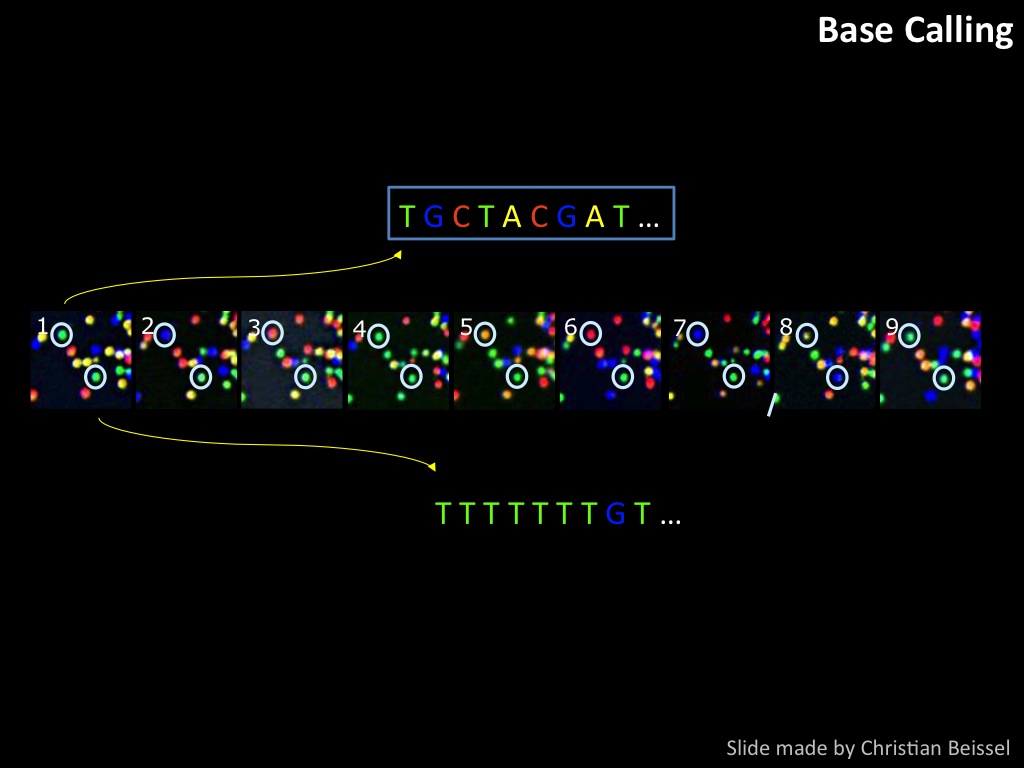
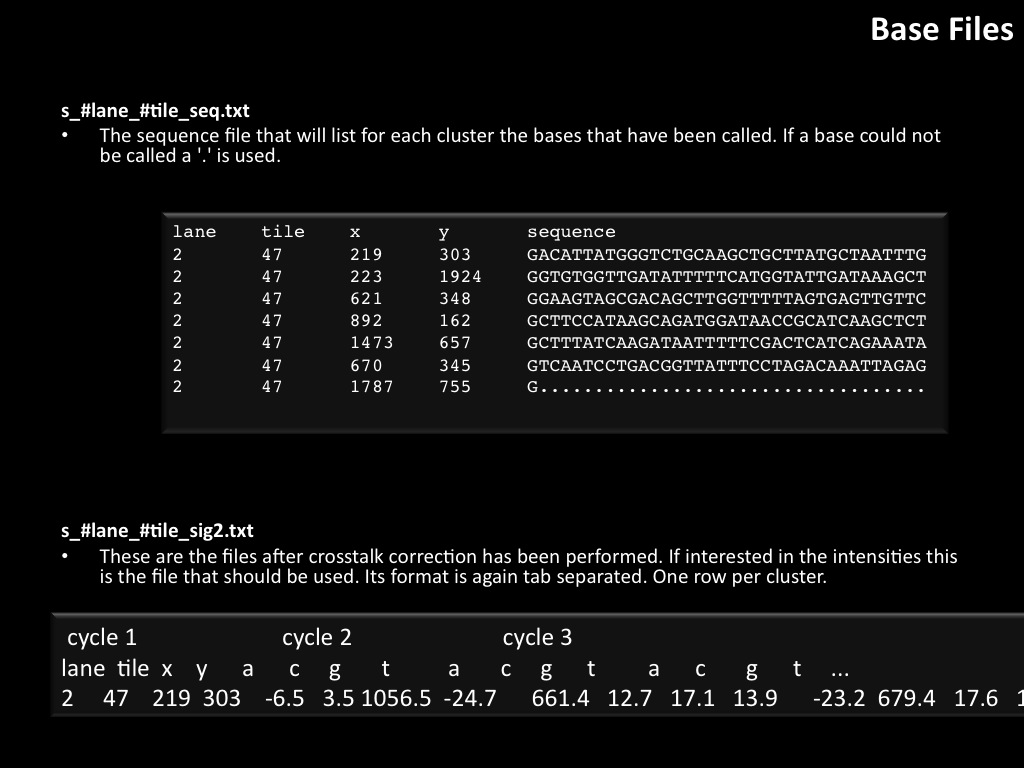
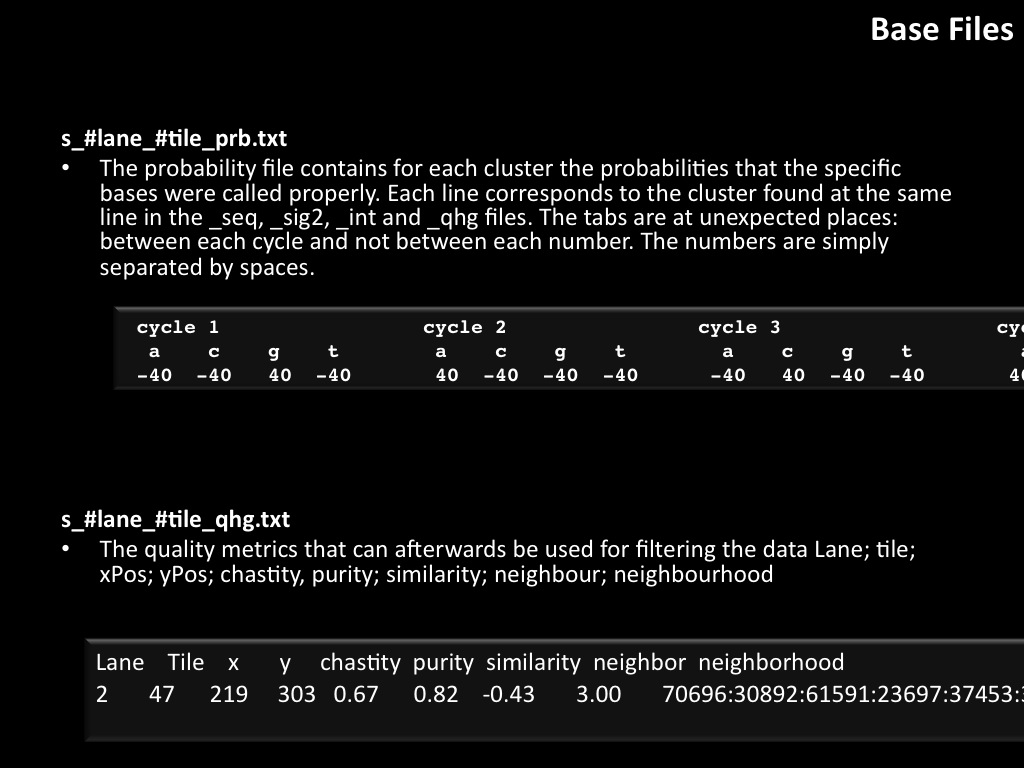
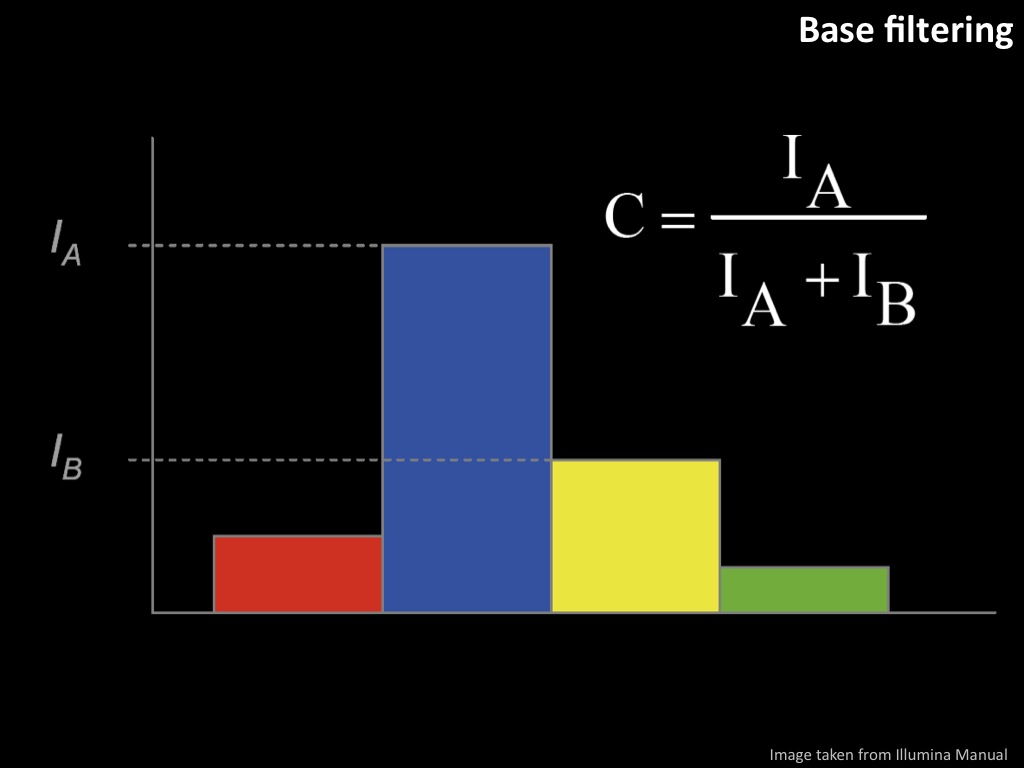
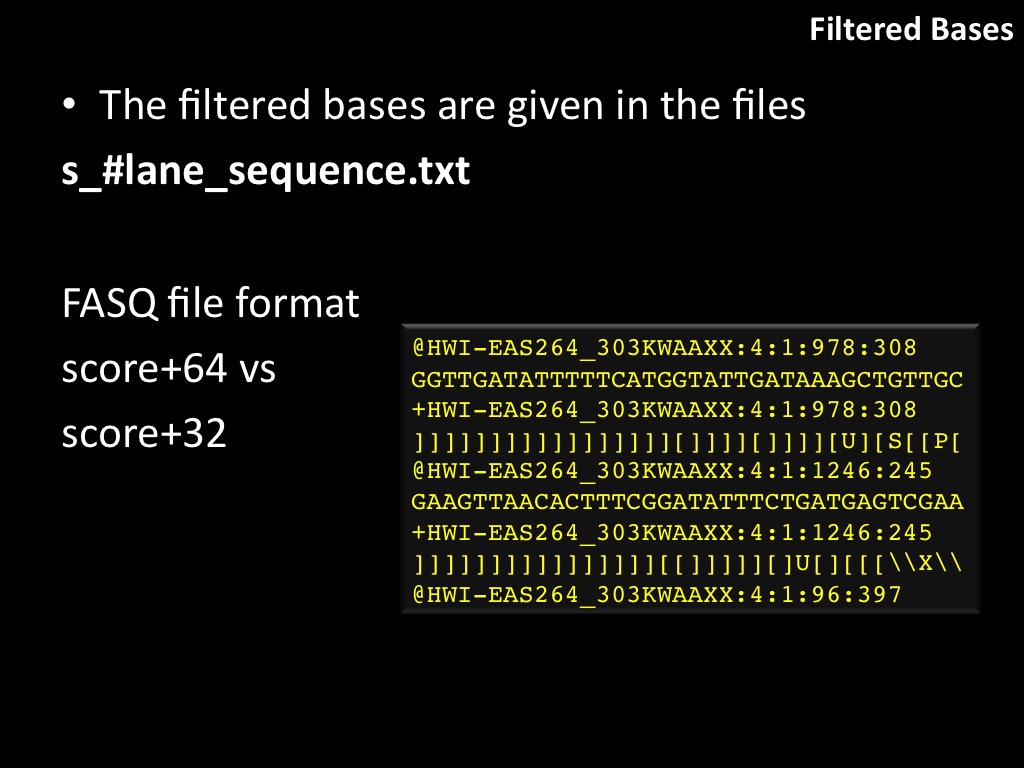
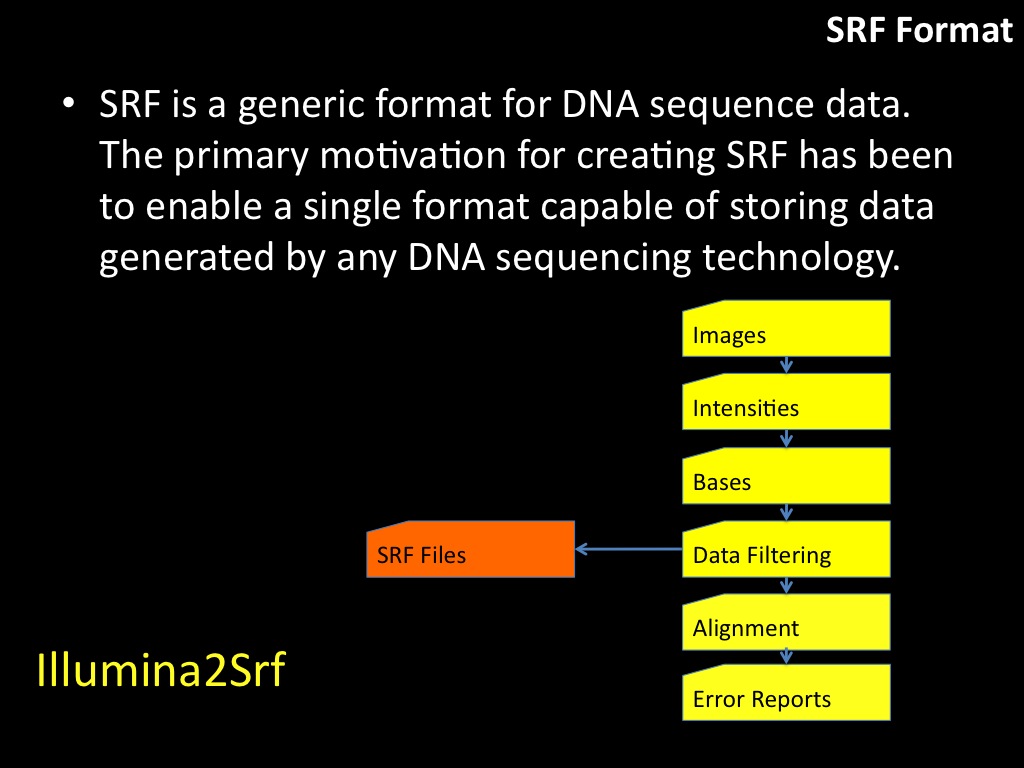
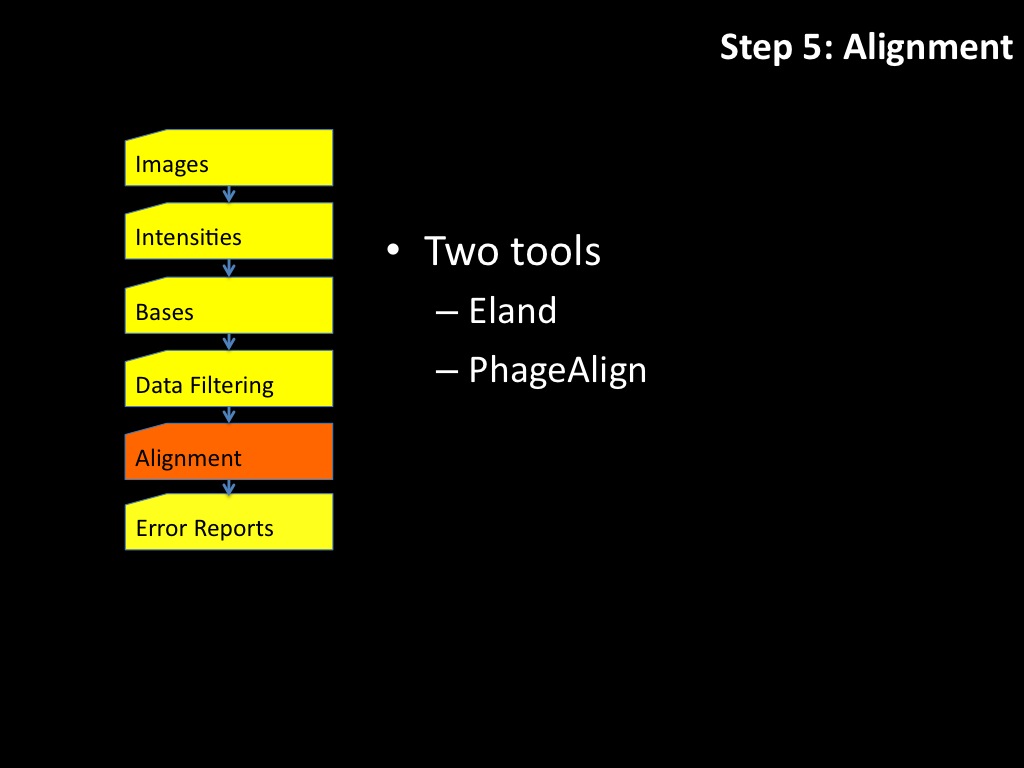
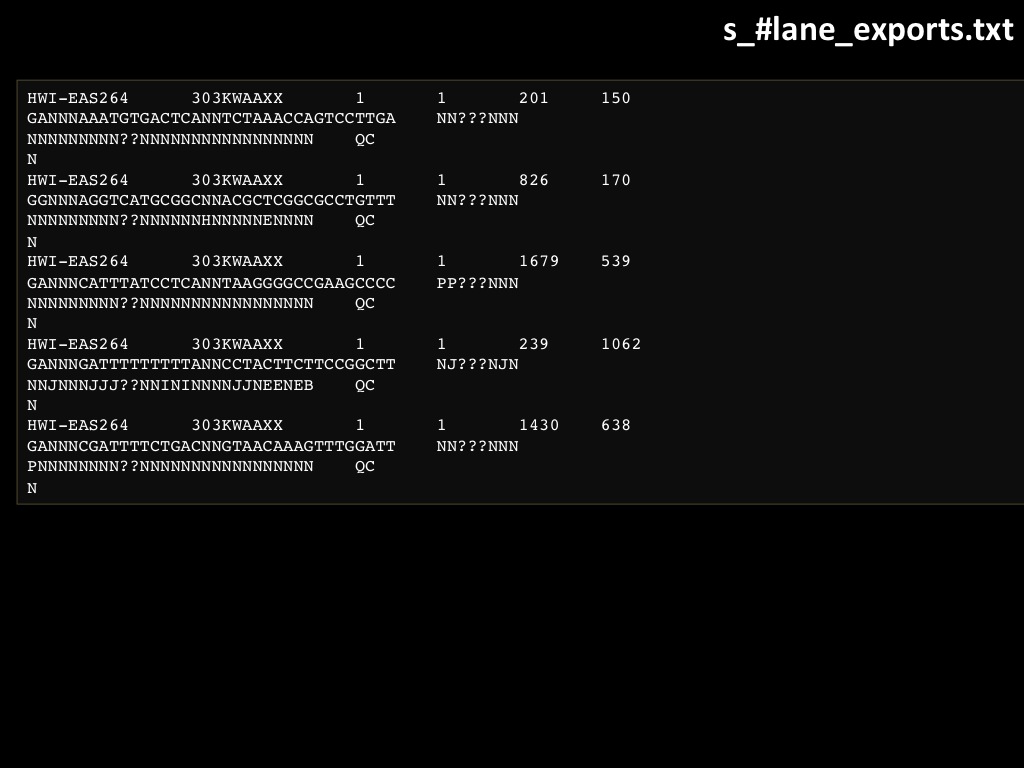
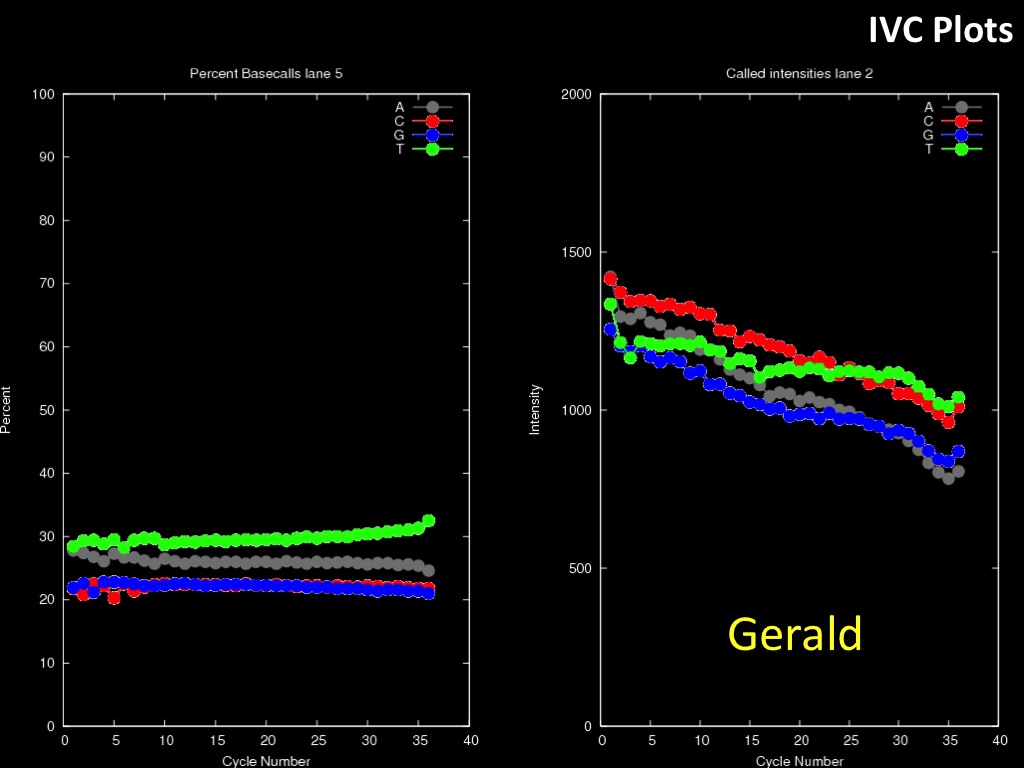
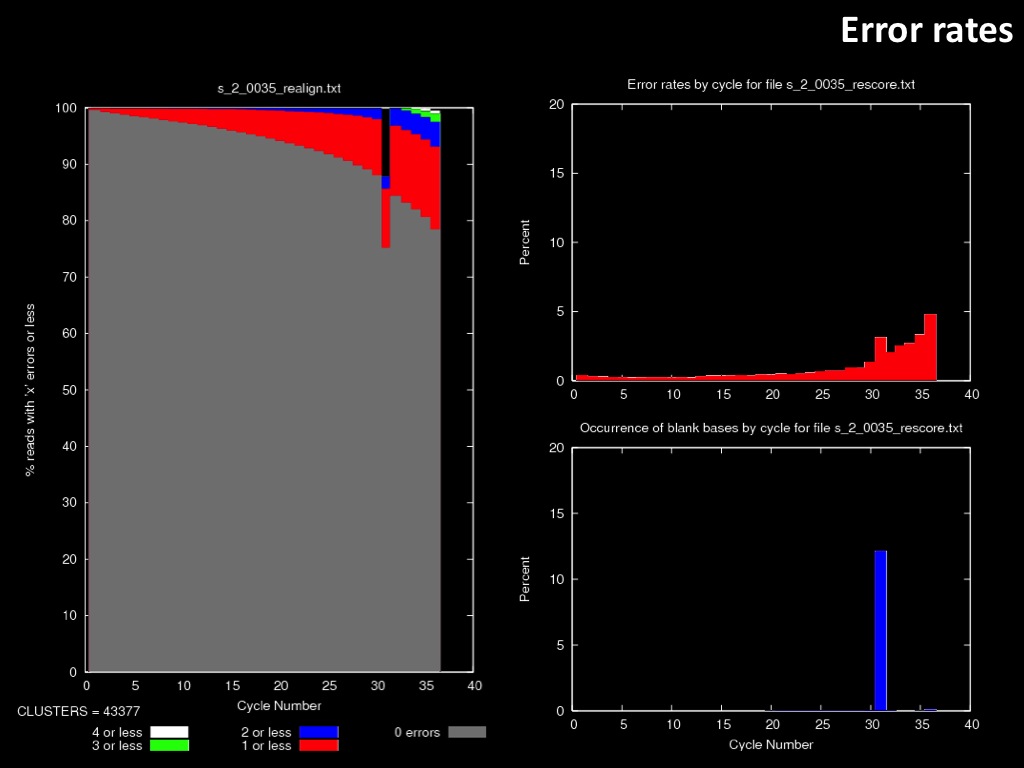
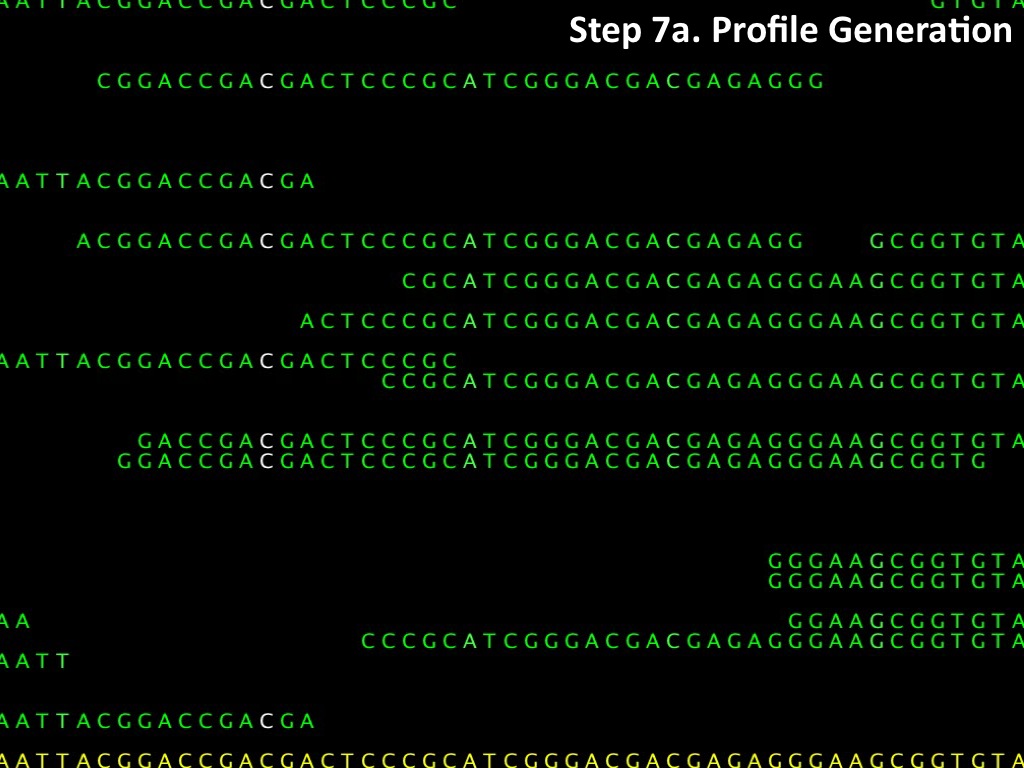
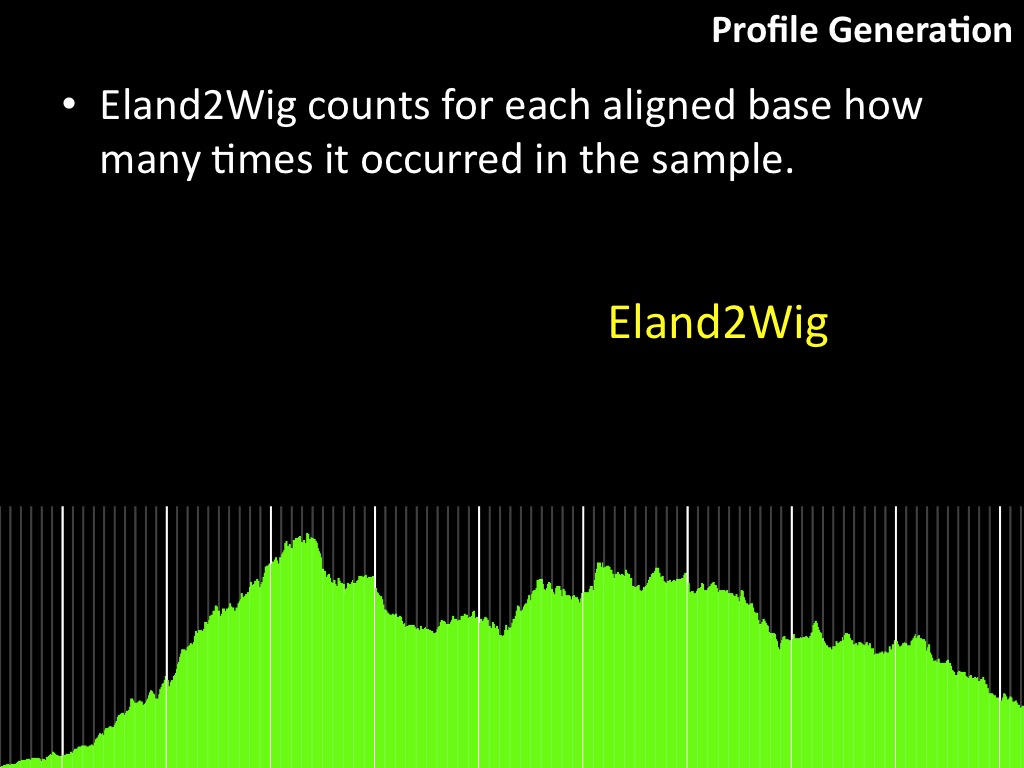
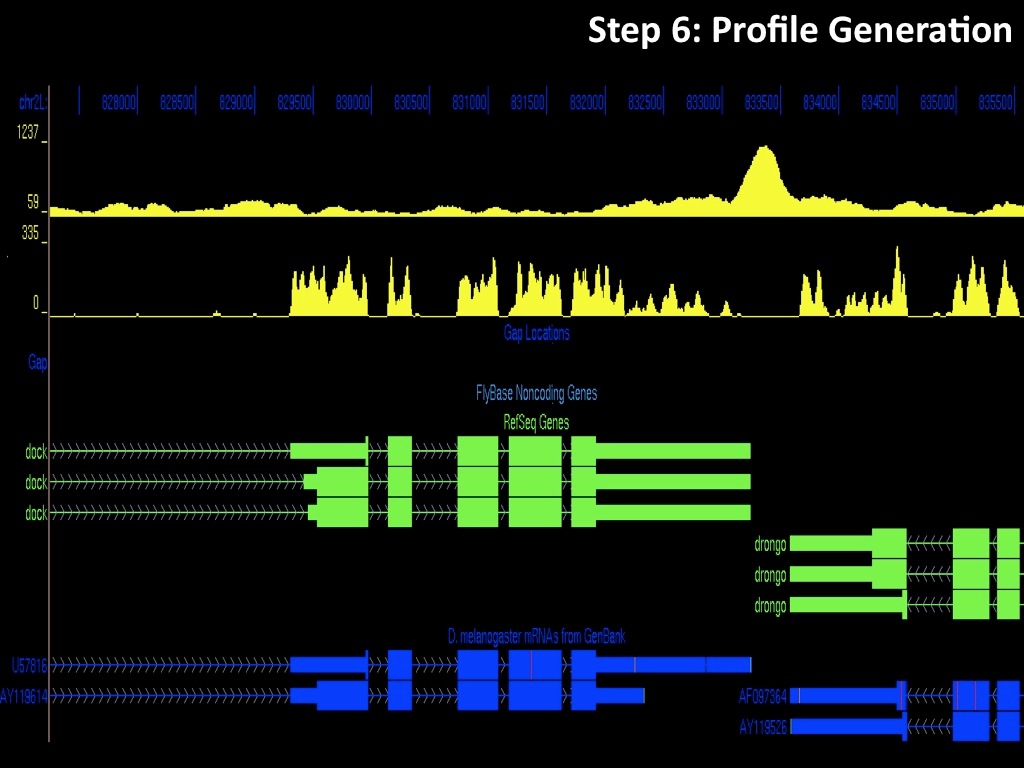
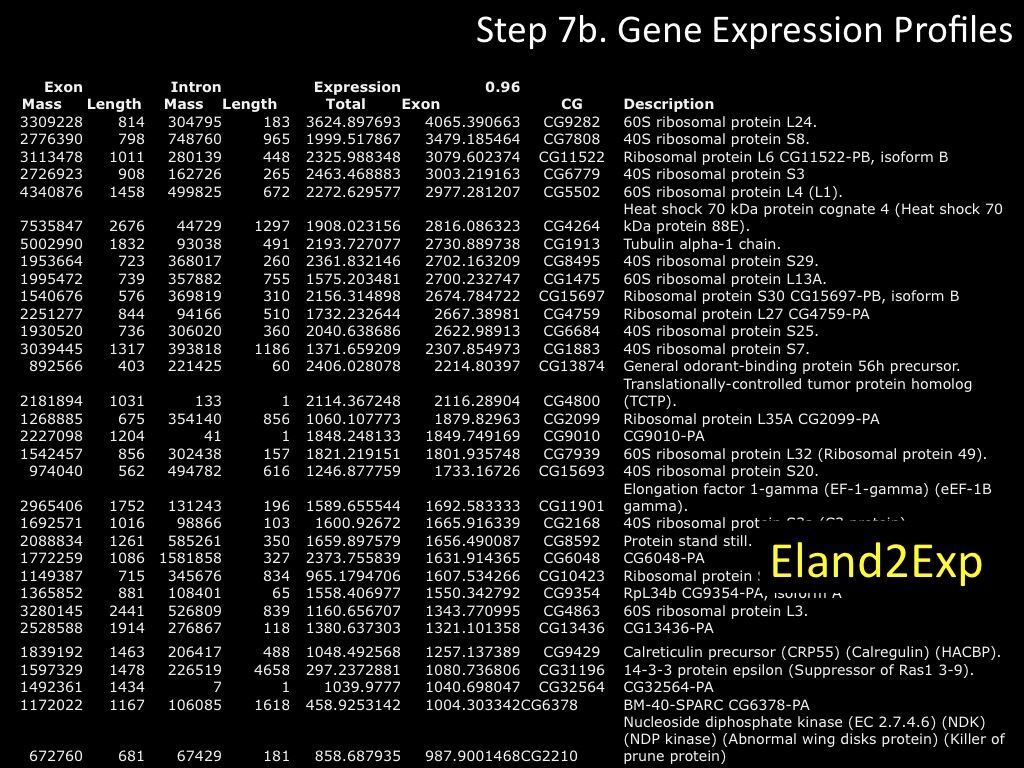
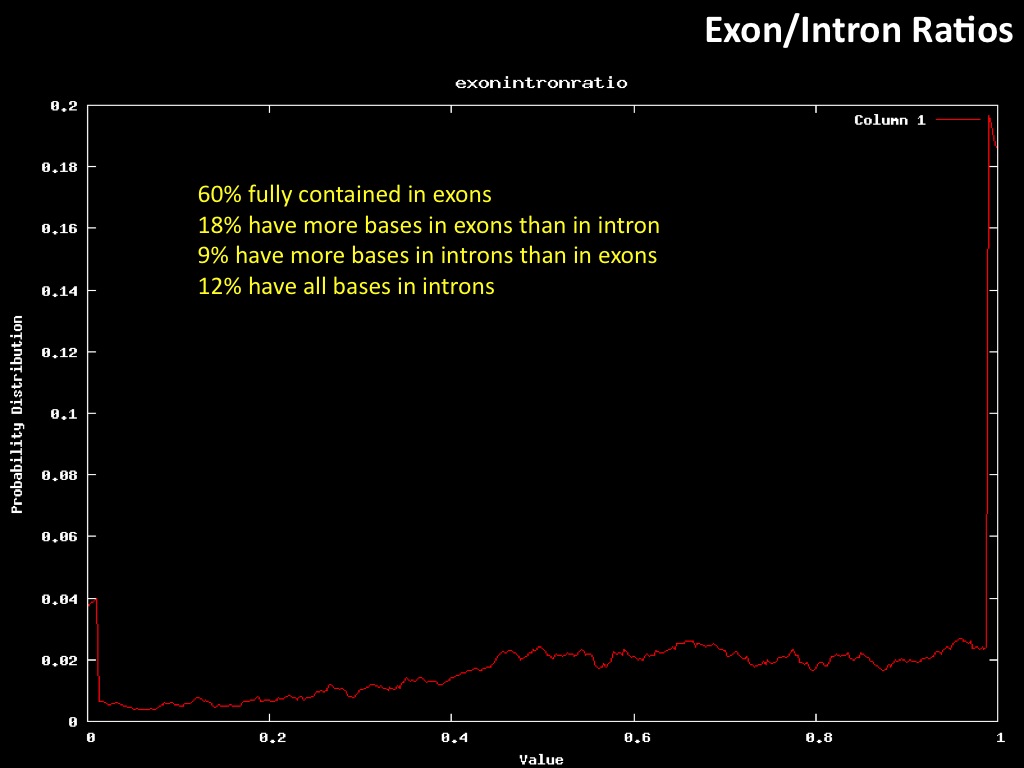
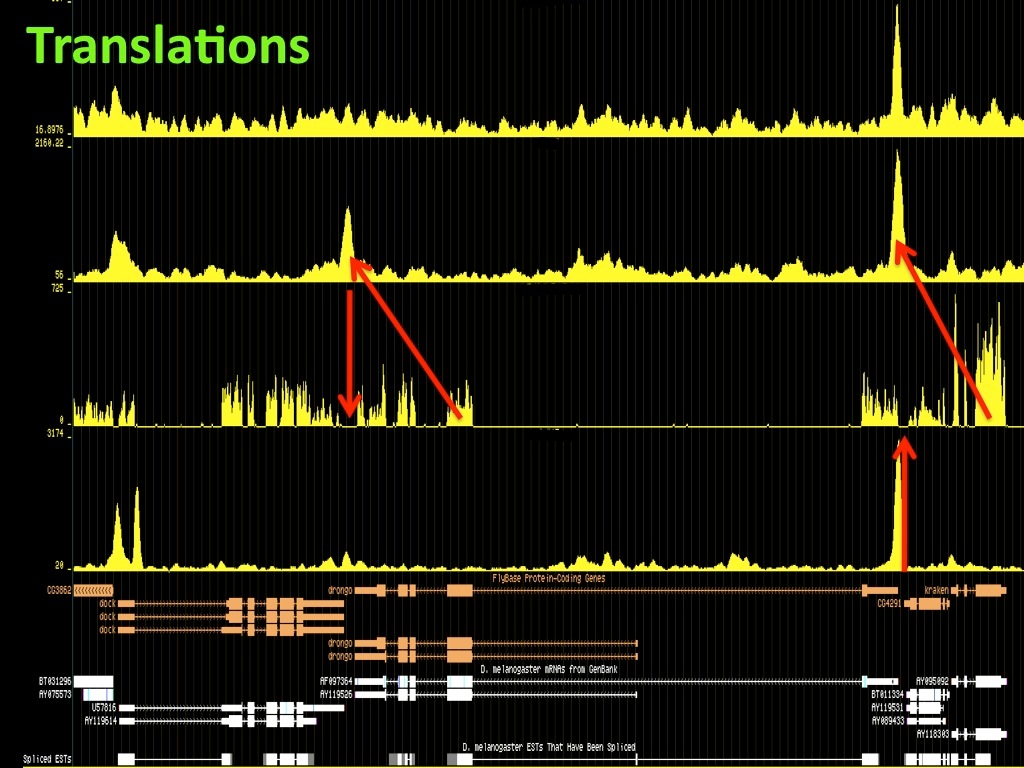
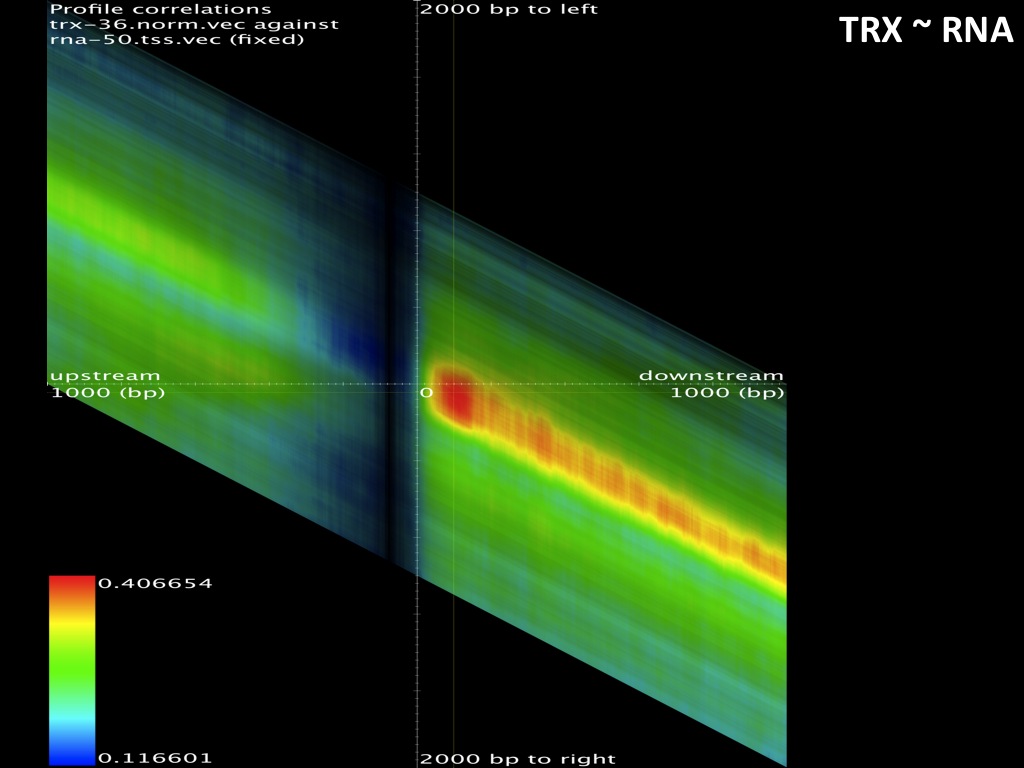
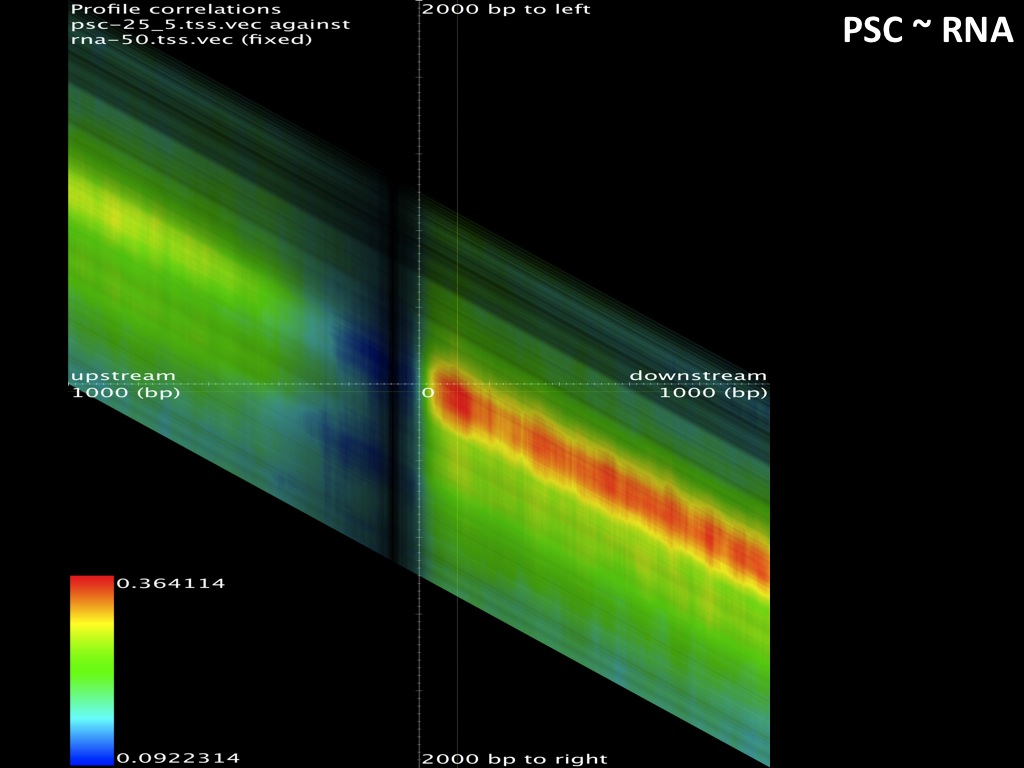
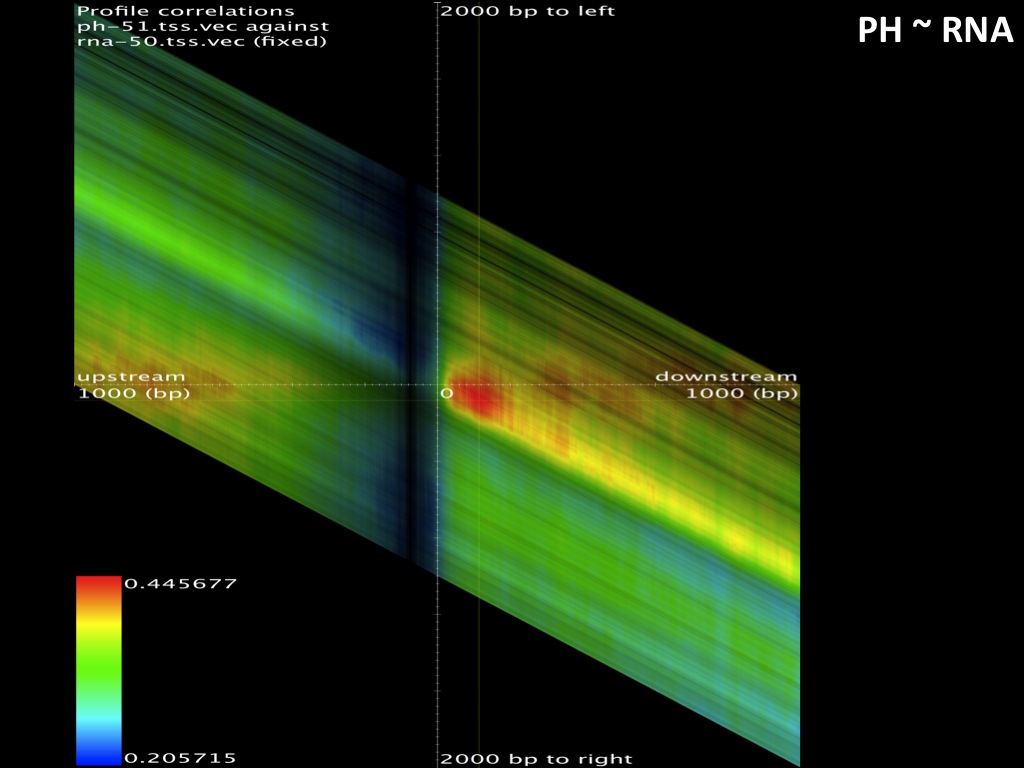
Abstract : I present a relative new technology: sequencing by synthesis (sometimes also called solexa sequencing, deep sequencing or digtal gene profiling). This technology has been booming the past few years because it can measure an entire genome expression profile at basepair resolution without predesigned probes. This makes it an excellent explorative tool and depending on the preprocessing of the sample, different information can be obtained. At the cost of a single micro-array one can measure methylation/acetylation patterns, genomic DNA, detect SNP's or measure the transcriptome. Never before has a single experiment, performed in 5 days, yielded 1.5Tb of data and 28'800'000'000 bases. Analyzing this data is a major challenge and the first part of the presentation covers the primary analysis. I will present the acquisitation of images, calculation and normalization of intensities, base calling, short fragment quality filtering and how one aligns 800'000'000 short fragments to a genome of choice. The second part of the presentation touches upon the problem of presenting this data to biologists and will illustrate our current approach where we translate a biological question to a signal processing question without resorting to arbitrary cutoff values. For this purpose, I will recycle some histone methylation and acetylation patterns that were previously published.
Keywords:
deep sequencing, sequencing by synthesis correlation analysis
Reference:
Werner Van Belle; Sequencing by synthesis: from samples to data patterns.; Presented at Novartis; Quantitative Biology Meetings; Switzerland; November 2008
Files: alignment2profile.avi
See also:
The research article.
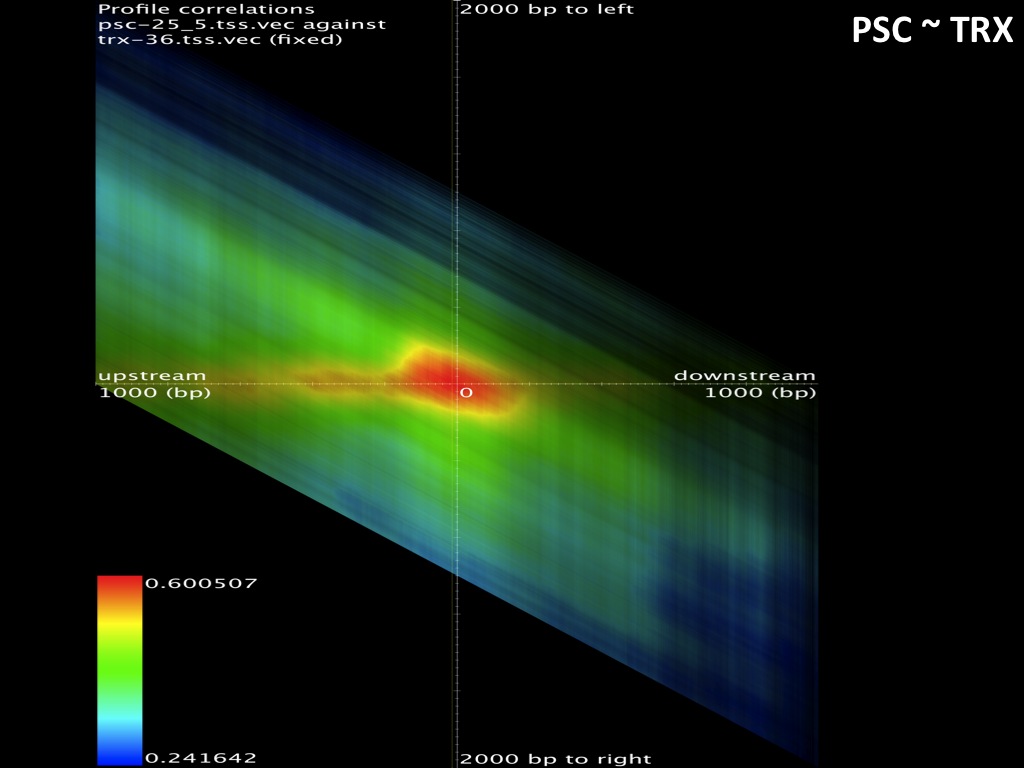
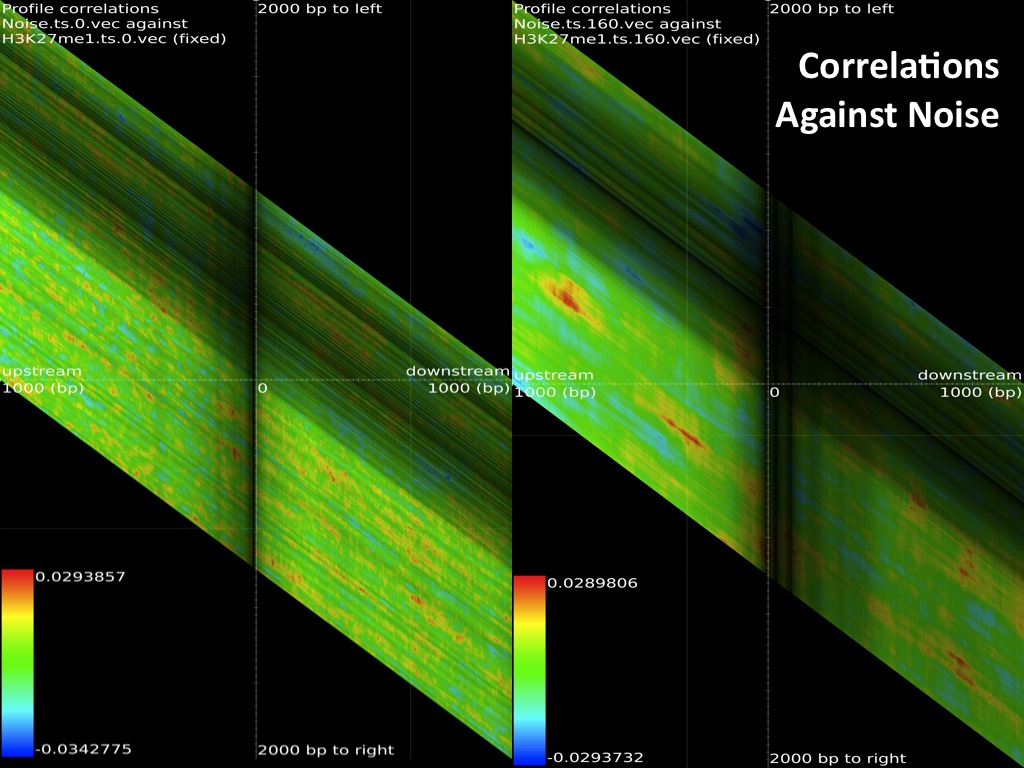
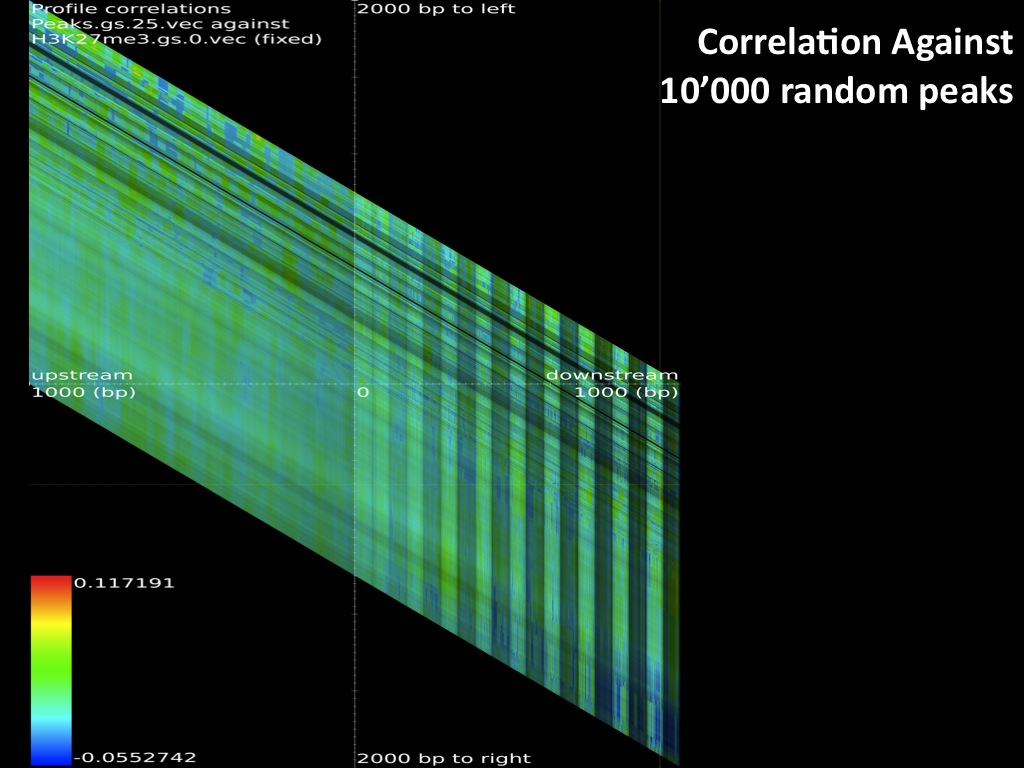
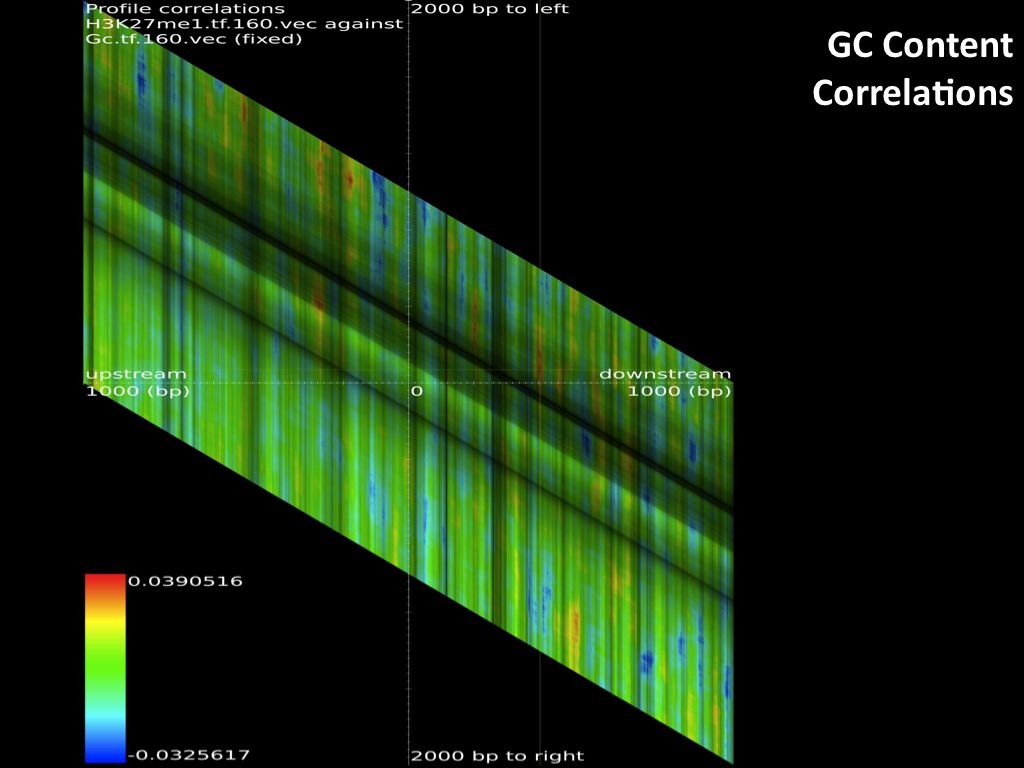
The browsable correlation maps between Histone Methylation and Acetylation tracks
Some posters [1, 2, 3]
Presentations of this research at FMI, Novartis, UNIL, University Zürich and ETH Zürich [1, 2]

















| http://werner.yellowcouch.org/ werner@yellowcouch.org |  |